 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Seita.9G510100.1.p | ||||||||
| Common Name | SETIT_034076mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1035aa MW: 115162 Da PI: 5.3218 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.1 | 1.1e-55 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenp 92
ke ++rwl++ ei++iL+n++++++t e+++rp+sgsl+L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+++vl+cyYah+een+
Seita.9G510100.1.p 20 KEaQNRWLRPAEICEILKNYRNFRITPEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKSGSIDVLHCYYAHGEENE 110
5559*************************************************************************************** PP
CG-1 93 tfqrrcywlLeeelekivlvhylevk 118
+fqrr+yw+Lee++ +ivlvhylevk
Seita.9G510100.1.p 111 NFQRRSYWMLEEDFMHIVLVHYLEVK 136
***********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 83.172 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.1E-77 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.3E-49 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| CDD | cd00102 | 2.60E-5 | 445 | 532 | No hit | No description |
| Gene3D | G3DSA:2.60.40.10 | 4.8E-8 | 445 | 530 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 9.33E-19 | 446 | 531 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 5.6E-9 | 448 | 525 | IPR002909 | IPT domain |
| CDD | cd00204 | 1.63E-12 | 626 | 738 | No hit | No description |
| SuperFamily | SSF48403 | 4.97E-16 | 637 | 741 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.3E-16 | 640 | 744 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.131 | 646 | 750 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.576 | 646 | 678 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.736 | 679 | 711 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.013 | 679 | 708 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3500 | 718 | 747 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.91E-8 | 851 | 903 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.07 | 852 | 874 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.114 | 853 | 882 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0029 | 854 | 873 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0019 | 875 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.542 | 876 | 900 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.6E-4 | 878 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1035 aa Download sequence Send to blast |
MAEARRYAIA PQLDIEQILK EAQNRWLRPA EICEILKNYR NFRITPEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHER LKSGSIDVLH CYYAHGEENE NFQRRSYWML 120 EEDFMHIVLV HYLEVKGGKS SSRIRGHDDM LQAARTDSPL SQLPSQTTEG ESSLSGQASE 180 YEETESDIYS GGAGYHPFSW MQQHENGGGP VMGASILSSY IPAPSVGNHQ GFLATTTTTD 240 FYSHGQDALP VALNEPGFGI AFDEADNQLD PSSLNGLVKP DQGVHRMAPP QIADPSKQFP 300 FTEGSGIESF TFDEVYSNGL SIKDADTVGT DEESLWQLPG AISSFPTEDS SQQNGRSLEE 360 AINHPLLKTQ SSSLSDILKD SFKKNDSFTR WMSKELGEVD DSPIKSSSGV YWNSEDTDNI 420 IEASSRDQLD QFTVDPVVAQ DQLFSIFDFS PSWAYAGSKT RVLITGRFLN SDEVQRCKWS 480 CMFGEVEVPA EISADGTLRC YSPSHKPGRV PFYVTCSNRL ACSEIREFEF RPSNSQHIDG 540 PTPHDIANKT YLQMRLDDLL SLGQDEYQAT VSNPTKDMID LSKKISSLMT DNDSWSELLK 600 LAGDNELATD DKQDQFFENR VKEKLHIWLV HKAGDGGKGP SVLDEEGQGV LHLAAALGYD 660 WAIRPTISAG VSINFRDAHG WTALHWAAFC GRERTVVALI ALGAAPGALT DPTPDFPTGS 720 TPADLASANG YKGISGFLAE SSLTSHLQTL NLKEAMWSNA PEISGLPGIG DVTERKLSPL 780 AGEGLLAGSM GDSLGAVRNA TQAAARIYQV FRMQSFQRKQ VVQYEDDNGA ISDDCALSLL 840 SVKPSKPGQL DPLHAAATRI QNKYRGWKGR KEFLLIRQRI IKIQAHVRGH QVRKHYRKII 900 WSVGIVEKVI LRWRRRGAGL RGFRSTEGAA EGTSSSGSDL IQNKPAEDDY DFLQQGRKQT 960 EERLQKALAR VKSMVQYPDA RDQYQRILNV VTKMQESQAM QEKMLESSTD MDEGLAMSDF 1020 EELWGDDMPM PGYI* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
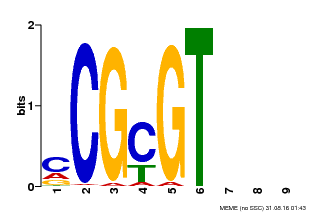
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Seita.9G510100.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004985393.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X2 | ||||
| TrEMBL | A0A368SUT7 | 0.0 | A0A368SUT7_SETIT; Uncharacterized protein | ||||
| STRING | Si034076m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Seita.9G510100.1.p |



