 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Seita.5G420700.1.p | ||||||||
| Common Name | SETIT_000062mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1331aa MW: 146935 Da PI: 8.7552 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 14 | 0.00014 | 1296 | 1322 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+Cp Cg++F+ s++ rH r+ H
Seita.5G420700.1.p 1296 YVCPepECGQTFRFVSDFSRHKRRtgH 1322
99********************98666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 4.7E-15 | 18 | 59 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.73 | 19 | 60 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 5.2E-13 | 20 | 53 | IPR003349 | JmjN domain |
| SMART | SM00558 | 2.1E-46 | 184 | 353 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 34.838 | 187 | 353 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 2.42E-24 | 200 | 350 | No hit | No description |
| Pfam | PF02373 | 3.5E-36 | 217 | 335 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 8.0E-4 | 1211 | 1235 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 15 | 1213 | 1235 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.907 | 1236 | 1265 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.5 | 1236 | 1260 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1238 | 1260 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.36E-14 | 1247 | 1300 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-9 | 1264 | 1290 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 9.7E-4 | 1266 | 1290 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.718 | 1266 | 1295 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1268 | 1290 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.8E-10 | 1291 | 1318 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.74 | 1296 | 1322 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.845 | 1296 | 1327 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1298 | 1322 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1331 aa Download sequence Send to blast |
MPSVQGELVP PWLKSLPLAP EFRPTASEFA DPIAYLLKIE PAAAPFGICK VVPPLPPPPK 60 RTTLGNLSRS FAALHPGDPS PTFPTRHQEL GLCPRRPRPA LKPVWHSPRR YTLPQFEAEA 120 GASRKALLAR LDVPPSRHLS PLDVEALFWR SSADRPVAVE YASDMPGSGF APCDARPTQL 180 PAANVGETAW NMRGVARSPA SLLRFLREEV PGVTSPMLYV AMLFSWFAWH VEDHDLHSLN 240 YLHSGAPKTW YGVPRDAALA FEDVVRVHGY GGEVNPLETF AMLGDKTTVM SPEVLVRSGI 300 PCCRLVQNAG EFVVTFPGSY HSGFSHGFNY GEASNIATPE WLRAAKEAAV RRASINRPPM 360 VSHYQLLYEL ALSMCLRDPS GGAMEPRSSR LKEKKKGEGE QLVKKIFVRN VIEDNKLLNH 420 FLSDGSSCII LPTSSNNGSA LSTLLSKSQS TTSRVSDVQC SSTETPKDSG HLPMNGALGK 480 NGELSSSKEI SASVCSGKEV PPTACMHDCV NMPGSLDANN AESDKGDVNN ADGILDQGLL 540 SCVTCGILSF SCVAVIKPRE CAAKWLMSAD SSLINKQLAG SGESHLIDAL QGSDFEMNRN 600 RIISDAASLD RNSALDLLAS AYGDASDSDE DVLNKKIQAS NVSNELISHT IESPPNSSSN 660 GGCDGTNMSS SSKERQQGPS SQSSQCIGNT NNGPKGVRTR NKYQLKMVLS EGFLPKDIYS 720 EMQKKVQCEP SRSNMTSTEP IHGTDCQASR NSATVCMDGN RSTTTTVDNL ATSIVKPDKD 780 SSRMHVFCLE HAIEVEKQLR TIGGAHIFLL CRPEYPKIEV EAKLLAEEME VKYDWKDIVF 840 KEASIEDRKK IQEVVQDEET IPTHSDWAVK LGINLYYSAN LAKSPLYNKQ LPYNRVIYKA 900 FGCSSPNNSP AKLKTYARRQ GRAKKIVLAG RWCGKVWMSN QVHPFLAHRI ESHEPEEIDE 960 IWSCYEKSNA DHVEHSSREA TSPRKSSSRA IEEKTSNREK EPLEKASIKK PKYIEEDNSE 1020 ALESAEKASA GKSNCRTSVE KMGKRKKELA EKANTKKLKH TEEDNSKALT GASEASPPLP 1080 SGMVVRSSSR IANRKNMLKS KMEEEDNGPA SHPKAKVEED SNDPAICSSA RSLRQNINVK 1140 KQTKKSRAEK RKAPSSAALK DEEQISDVKG FSVTKQQLSS HKQKNKVEET QQMKKTRERK 1200 GAPPSSPKHG EEYACDIEGC SMSFGTKQEL SLHKRDICPV QGCRRKFFSH KYLLQHRKVH 1260 NDDRPLKCSW KGCDMAFKWP WARTEHMRVH TGDRPYVCPE PECGQTFRFV SDFSRHKRRT 1320 GHAAKVKAKK * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 1e-70 | 12 | 374 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 1e-70 | 12 | 374 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1043 | 1058 | KRKKELAEKANTKKLK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3 with a specific activity for H3K27me3 and H3K27me2. No activity on H3K4me3, H3K9me3, H3K27me1 and H3K36me3. Involved in biotic stress response. May demethylate H3K27me3-marked defense-related genes and increase their basal and induced expression levels during pathogen infection. {ECO:0000269|PubMed:24280387}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
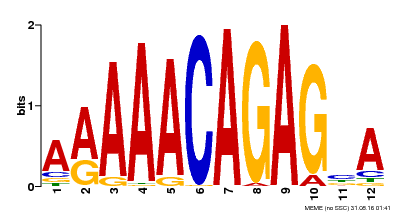
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Seita.5G420700.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt stress, abscisic acid (ABA), jasmonic acid (JA), the ethylene precursor ACC and infection by the bacterial pathogen Xanthomonas oryzae pv. oryzae. {ECO:0000269|PubMed:24280387}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004970976.1 | 0.0 | lysine-specific demethylase JMJ705 | ||||
| Swissprot | Q5N712 | 0.0 | JM705_ORYSJ; Lysine-specific demethylase JMJ705 | ||||
| TrEMBL | A0A368RF07 | 0.0 | A0A368RF07_SETIT; Uncharacterized protein | ||||
| STRING | Si000062m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5346 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Seita.5G420700.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



