 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676782592 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 426aa MW: 46675.3 Da PI: 10.215 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 41.5 | 2.9e-13 | 347 | 399 | 5 | 57 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkeva 57
+r++r++kNRe+A rsR+RK+a++ eLe ++++L++ N++L+ + e+ k+ +
676782592 347 RRQKRMIKNRESAARSRARKQAYTLELEAEIENLKQLNQDLQRKQAEIMKTQK 399
79***************************************877666666544 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.4E-12 | 343 | 408 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.461 | 345 | 396 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 2.25E-9 | 347 | 396 | No hit | No description |
| Pfam | PF00170 | 4.3E-11 | 347 | 400 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 2.2E-12 | 347 | 404 | No hit | No description |
| CDD | cd14707 | 6.90E-24 | 347 | 399 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 350 | 365 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 426 aa Download sequence Send to blast |
MAVRVEVKGK KGLIRFVFKK VMGTHINFNN LDGDDSGGNG SNNNQSKPPL ARQSSLYSLT 60 FDELQSTLGE PGKDFGSMNM DELLKNIWSA EETQAIMTTT SSIAAVQPSS GCVVPGGNLL 120 QRQGSLTLPR TLSQKTVDEV WKNLVSKESC NGNSGTDAPE RQQTLGEMTL EDFLLRAGVV 180 KEDFDSNNQN SSLGFYANNG ASAALGFGFG QPNQNNISFN GNNNSMILNQ APGLGVKVGG 240 TMQQQHHHQP HQQQQLQQPH QRLPPTIYPK QANVTFAAPV NIVKKGLYEA PANNNSGFAT 300 MGGGGVTVAS TSPGTSSAEN NAWSSPVPYV FGRGRRSNTG LEKVVERRQK RMIKNRESAA 360 RSRARKQAYT LELEAEIENL KQLNQDLQRK QAEIMKTQKN ELKETSKQRP WLAKTQCLRR 420 TLTGPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Could participate in abscisic acid-regulated gene expression. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00087 | SELEX | Transfer from AT1G49720 | Download |
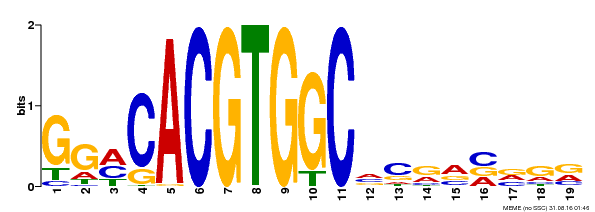
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676782592 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by abscisic acid (ABA) and cold. {ECO:0000269|PubMed:10636868}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001322564.1 | 0.0 | abscisic acid responsive element-binding factor 1 | ||||
| Refseq | NP_564551.1 | 0.0 | abscisic acid responsive element-binding factor 1 | ||||
| Swissprot | Q9M7Q5 | 0.0 | AI5L4_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 4 | ||||
| TrEMBL | A0A178WH34 | 0.0 | A0A178WH34_ARATH; AtABF1 | ||||
| STRING | fgenesh2_kg.1__4055__AT1G49720.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM734 | 27 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49720.1 | 0.0 | abscisic acid responsive element-binding factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



