 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676718402 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 683aa MW: 75586.1 Da PI: 6.0173 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 97.2 | 1.4e-30 | 65 | 149 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+++e+laL+++r++m++++r++ lk+plWe+vs+k+ e g++rs+k+Ckek+en++k+yk++ke++++r++++ +++f+qlea
676718402 65 RWPREETLALLRIRSDMDSTFRDATLKAPLWEHVSRKLLELGYKRSAKKCKEKFENVQKYYKRTKETRGGRHDGK--AYKFFSQLEA 149
8*********************************************************************86555..5******985 PP
| |||||||
| 2 | trihelix | 103.8 | 1.3e-32 | 445 | 529 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
rW+k e+laLi++r+ me r++++ k+ lWee+s m++ g++r++k+Ckekwen+nk+ykk+ke++k+r +++ +tcpyf++l+
676718402 445 RWPKAEILALINLRSGMEPRYQDNVPKGLLWEEISTSMKRMGYNRNAKRCKEKWENINKYYKKVKESNKQR-PQDAKTCPYFHRLD 529
8*********************************************************************8.99999*******97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 6.946 | 58 | 122 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.0076 | 62 | 124 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 1.1E-20 | 64 | 150 | No hit | No description |
| CDD | cd12203 | 3.58E-27 | 64 | 129 | No hit | No description |
| PRINTS | PR01217 | 7.0E-10 | 153 | 165 | No hit | No description |
| PRINTS | PR01217 | 7.0E-10 | 194 | 206 | No hit | No description |
| PRINTS | PR01217 | 7.0E-10 | 206 | 227 | No hit | No description |
| PRINTS | PR01217 | 7.0E-10 | 362 | 387 | No hit | No description |
| PRINTS | PR01217 | 7.0E-10 | 400 | 412 | No hit | No description |
| SMART | SM00717 | 0.011 | 442 | 504 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 6.667 | 444 | 502 | IPR017877 | Myb-like domain |
| Pfam | PF13837 | 8.7E-22 | 444 | 529 | No hit | No description |
| CDD | cd12203 | 4.60E-30 | 445 | 509 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:0010090 | Biological Process | trichome morphogenesis | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0032876 | Biological Process | negative regulation of DNA endoreduplication | ||||
| GO:0042631 | Biological Process | cellular response to water deprivation | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:2000037 | Biological Process | regulation of stomatal complex patterning | ||||
| GO:2000038 | Biological Process | regulation of stomatal complex development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 683 aa Download sequence Send to blast |
MEQVGGGGGG GGGNEVVEEA SPISSRPPAN LEELMRFSAA TDDGGGLGGG GGAGGGGSSS 60 SSGNRWPREE TLALLRIRSD MDSTFRDATL KAPLWEHVSR KLLELGYKRS AKKCKEKFEN 120 VQKYYKRTKE TRGGRHDGKA YKFFSQLEAL NTTPPPSLPL HPPPSSSSLD VTPLSVANPI 180 QQQPPPSSQF PVFPLPQTQT PQPLQPHTVS FTPSPAPPPP MASTFPGVTF SSHSSSTASG 240 MGSDDDEDDM DVDQAGPSSR KRKRGNRGTG GGGKMMELFE GLVRQVMQKQ AAMQRSFLEA 300 LEKREQERLD REEAWKRQEM SRLAREHEIM SQERAASASR DAAIISLIQK ITGHTIQLPP 360 SFSSQPPQPS PPPHQPPPAA KRPSSQAVEP PQLQPIMAIP QQQVLPPPPA PHQPDHKQQQ 420 QQQQQQQPQQ EMIMSSDQSL PSSSRWPKAE ILALINLRSG MEPRYQDNVP KGLLWEEIST 480 SMKRMGYNRN AKRCKEKWEN INKYYKKVKE SNKQRPQDAK TCPYFHRLDL LYRNKVLGSG 540 GGSSASGLPQ DQISTVQKQS PVSAVKPAQE GVVDVHQPHG SASSEEEEPR EESPQGAEKP 600 EDLVMRELMQ QQQRQQHEDS MIGEYEKIEE SHNYNNMEEE EEDMDEEELD EDEKSAAYEI 660 AFQSPANRGG NGHTEPPFLT MVQ |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 107 | 115 | KRSAKKCKE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription repressor that binds specific DNA sequence such as GT3 box 5'-GGTAAA-3' in the SDD1 promoter. Negative regulator of water use efficiency (WUE) via the promotion of stomatal density and distribution by the transcription repression of SDD1. Regulates the expression of several cell cycle genes and endoreduplication, especially in trichomes where it prevents ploidy-dependent plant cell growth. {ECO:0000269|PubMed:19717615, ECO:0000269|PubMed:21169508}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00177 | DAP | Transfer from AT1G33240 | Download |
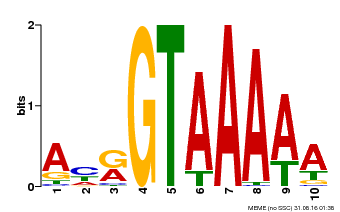
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676718402 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by water stress. {ECO:0000269|PubMed:21169508}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY140085 | 0.0 | AY140085.1 Arabidopsis thaliana DNA-binding factor, putative (At1g33240) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009123747.1 | 0.0 | PREDICTED: trihelix transcription factor GTL1 isoform X1 | ||||
| Refseq | XP_013717446.1 | 0.0 | trihelix transcription factor GTL1-like isoform X1 | ||||
| Swissprot | Q9C882 | 0.0 | GTL1_ARATH; Trihelix transcription factor GTL1 | ||||
| TrEMBL | A0A397Y2Y6 | 0.0 | A0A397Y2Y6_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4FFY6 | 0.0 | M4FFY6_BRARP; Uncharacterized protein | ||||
| STRING | Bra040010.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6226 | 27 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G33240.1 | 1e-150 | GT-2-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



