 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G33240.1 | ||||||||
| Common Name | ATGTL1, AT-GTL1, GTL1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 669aa MW: 74212.7 Da PI: 5.8532 | ||||||||
| Description | GT-2-like 1 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 96.4 | 2.6e-30 | 62 | 146 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+++e+laL+++r++m++++r++ lk+plWe+vs+k+ e g++rs+k+Ckek+en++k+yk++ke++++r++++ +++f+qlea
AT1G33240.1 62 RWPREETLALLRIRSDMDSTFRDATLKAPLWEHVSRKLLELGYKRSSKKCKEKFENVQKYYKRTKETRGGRHDGK--AYKFFSQLEA 146
8*********************************************************************86555..5******985 PP
| |||||||
| 2 | trihelix | 104.3 | 9.1e-33 | 435 | 519 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
rW+k e+laLi++r+ me r++++ k+ lWee+s m++ g++r++k+Ckekwen+nk+ykk+ke++kkr +++ +tcpyf++l+
AT1G33240.1 435 RWPKAEILALINLRSGMEPRYQDNVPKGLLWEEISTSMKRMGYNRNAKRCKEKWENINKYYKKVKESNKKR-PQDAKTCPYFHRLD 519
8*********************************************************************8.99999*******97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 6.934 | 55 | 119 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.015 | 59 | 121 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 5.04E-26 | 61 | 126 | No hit | No description |
| Pfam | PF13837 | 2.1E-20 | 61 | 147 | No hit | No description |
| SMART | SM00717 | 0.011 | 432 | 494 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 6.667 | 434 | 492 | IPR017877 | Myb-like domain |
| Pfam | PF13837 | 5.8E-22 | 434 | 519 | No hit | No description |
| CDD | cd12203 | 2.64E-29 | 435 | 499 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:0010090 | Biological Process | trichome morphogenesis | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0032876 | Biological Process | negative regulation of DNA endoreduplication | ||||
| GO:0042631 | Biological Process | cellular response to water deprivation | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:2000037 | Biological Process | regulation of stomatal complex patterning | ||||
| GO:2000038 | Biological Process | regulation of stomatal complex development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000014 | anatomy | rosette leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000282 | anatomy | trichome | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0000351 | anatomy | guard mother cell | ||||
| PO:0004011 | anatomy | initial cell | ||||
| PO:0006019 | anatomy | leaf abaxial epidermis | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 669 aa Download sequence Send to blast |
MEQGGGGGGN EVVEEASPIS SRPPANNLEE LMRFSAAADD GGLGGGGGGG GGGSASSSSG 60 NRWPREETLA LLRIRSDMDS TFRDATLKAP LWEHVSRKLL ELGYKRSSKK CKEKFENVQK 120 YYKRTKETRG GRHDGKAYKF FSQLEALNTT PPSSSLDVTP LSVANPILMP SSSSSPFPVF 180 SQPQPQTQTQ PPQTHNVSFT PTPPPLPLPS MGPIFTGVTF SSHSSSTASG MGSDDDDDDM 240 DVDQANIAGS SSRKRKRGNR GGGGKMMELF EGLVRQVMQK QAAMQRSFLE ALEKREQERL 300 DREEAWKRQE MARLAREHEV MSQERAASAS RDAAIISLIQ KITGHTIQLP PSLSSQPPPP 360 YQPPPAVTKR VAEPPLSTAQ SQSQQPIMAI PQQQILPPPP PSHPHAHQPE QKQQQQPQQE 420 MVMSSEQSSL PSSSRWPKAE ILALINLRSG MEPRYQDNVP KGLLWEEIST SMKRMGYNRN 480 AKRCKEKWEN INKYYKKVKE SNKKRPQDAK TCPYFHRLDL LYRNKVLGSG GGSSTSGLPQ 540 DQKQSPVTAM KPPQEGLVNV QQTHGSASTE EEEPIEESPQ GTEKPEDLVM RELIQQQQQL 600 QQQESMIGEY EKIEESHNYN NMEEEEDQEM DEEELDEDEK SAAFEIAFQS PANRGGNGHT 660 EPPFLTMVQ |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 104 | 112 | KRSSKKCKE |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.28206 | 0.0 | flower| root| seed| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 42562478 | 0.0 | ||||
| Genevisible | 261594_at | 0.0 | ||||
| Expression Atlas | AT1G33240 | - | ||||
| AtGenExpress | AT1G33240 | - | ||||
| ATTED-II | AT1G33240 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Mostly expressed in siliques, and, to a lower extent, in growing root hairs, leaves, stems, and flowers. Present in abaxial epidermal cells, predominantly in guard cells, pavement cells, and meristemoids. {ECO:0000269|PubMed:21169508, ECO:0000269|PubMed:9501260}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a plant transcriptional activator that contains two separate, but similar, trihelix DNA-binding domains, similar to GT-2. Gene is expressed in all aerial parts of the plant, with higher level of expression in siliques. At-GTL2 was thought to be a duplicated copy of this gene but is likely to be a cloning artefact, the result of a chimeric clone. Regulates ploidy-dependent cell growth in trichome. | |||||
| UniProt | Transcription repressor that binds specific DNA sequence such as GT3 box 5'-GGTAAA-3' in the SDD1 promoter. Negative regulator of water use efficiency (WUE) via the promotion of stomatal density and distribution by the transcription repression of SDD1. Regulates the expression of several cell cycle genes and endoreduplication, especially in trichomes where it prevents ploidy-dependent plant cell growth. {ECO:0000269|PubMed:19717615, ECO:0000269|PubMed:21169508}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
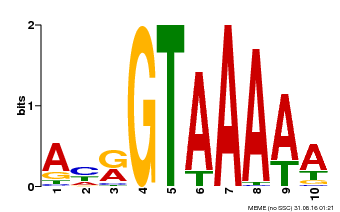
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00177 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G33240.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by water stress. {ECO:0000269|PubMed:21169508}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- ATRM (Manually Curated Target Genes) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Target Gene (A: Activate/R: Repress) | |||||
| ATRM | AT1G04110(R) | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G33240 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC021045 | 0.0 | AC021045.2 Arabidopsis thaliana chromosome I BAC T9L6 genomic sequence, complete sequence. | |||
| GenBank | AC027035 | 0.0 | AC027035.5 Arabidopsis thaliana chromosome 1 BAC T16O9 genomic sequence, complete sequence. | |||
| GenBank | AJ003215 | 0.0 | AJ003215.1 Arabidopsis thaliana GTL1 gene. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_174594.1 | 0.0 | GT-2-like 1 | ||||
| Swissprot | Q9C882 | 0.0 | GTL1_ARATH; Trihelix transcription factor GTL1 | ||||
| TrEMBL | A0A1P8ATZ4 | 0.0 | A0A1P8ATZ4_ARATH; GT-2-like 1 | ||||
| STRING | AT1G33240.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6226 | 27 | 46 | Representative plant | OGRP663 | 15 | 73 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G33240.1 |
| Entrez Gene | 840218 |
| iHOP | AT1G33240 |
| wikigenes | AT1G33240 |



