 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_011072923.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Pedaliaceae; Sesamum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1297aa MW: 144320 Da PI: 8.5848 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.3 | 0.00025 | 1206 | 1228 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
XP_011072923.1 1206 CPvkGCGKKFFSHKYLVQHRRVH 1228
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 12.5 | 0.00046 | 1264 | 1290 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ Cg++F+ s++ rH r+ H
XP_011072923.1 1264 YVCTeaGCGQTFRFVSDFSRHKRKtgH 1290
99********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 4.3E-15 | 3 | 44 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.966 | 4 | 45 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 1.4E-14 | 5 | 38 | IPR003349 | JmjN domain |
| SMART | SM00558 | 6.0E-53 | 168 | 337 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.91 | 168 | 337 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.81E-26 | 186 | 342 | No hit | No description |
| Pfam | PF02373 | 6.8E-38 | 201 | 320 | IPR003347 | JmjC domain |
| SMART | SM00355 | 20 | 1181 | 1203 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.445 | 1204 | 1233 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 1204 | 1228 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.8E-5 | 1205 | 1232 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1206 | 1228 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.74E-9 | 1220 | 1262 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-8 | 1233 | 1258 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1234 | 1263 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1234 | 1258 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1236 | 1258 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 8.8E-8 | 1252 | 1286 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-9 | 1259 | 1287 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.344 | 1264 | 1295 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.19 | 1264 | 1290 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1266 | 1290 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1297 aa Download sequence Send to blast |
MPVAPEYHPT LAEFQDPIAY IFKIEKEASK YGICKIVPPV PAASKKTVIA NLNKSLLARS 60 TDPESGPTFT TRQQQIGFCP RKHRPVQKPV WQSGEKYTLT EFEVKAKNFE KIYLKKYAKK 120 GLNALEMETV YWNATVDKPF QVEYANDMPG SAFVAQKGCG KKNESTITVG ETEWNMRRVS 180 RENLSLLRFM KEEIPGVTSP MVYVAMMFSW FAWHVEDHDL HSLNYLHMGA GKTWYGVPRE 240 AAVAFEEVIR EHGYGGEINP LVTFATLGEK TTVMSPEVLL SAGVPCCRLV QNAGEFVVTF 300 PRAYHSGFSH GFNCGEAANI ATPEWLRFAK EAAIRRAAIN CPPMVSHFQL LYDLALSLCS 360 GVPKSIAMGP RSSRLKDRKK GEGEMLIKEL FLQDMMQNND MLHSLGKGSS IVLLPQNSLS 420 HSIYNNTSSG FQSTAKSRLF PSLCSPDLEL KTASYDARDE FLLDRKHGIK QPRGHAVNRK 480 SVSSLCSSSE VPSMAPCAEQ IDSEIKRASQ HEQGLFSCVT CGILCFACAA IVQPTEAAAH 540 YLMSADCSKF SYWGTGDDDH NHTRHAKSPN TNLGSGLVLR KKHGPSDVPI SADQIRSVNG 600 ESVGVVSNSK AHKEPSSLGL LALTYGNSSD SEEEETEADL PVDGCGTSKS DSPEDGHACD 660 NIDSKLNCRK EMSSQISDSN AMFGPPIAKC NGGDPQSSNC SDEFPTDNCT VVESNSCTHR 720 SRHRTKSRHD TSNSLTHKTE ATVSTGLTPL EDKNTPFPAR SDEDSYRLHV FCLQHALQVE 780 KRLSQIGGAN VFLVCHPDFP KLESQAKKVA EELESDWLWS EISFREATEE DEEIMRLALE 840 SENAIHGNGD WAVKMGINLF FSANLSRSPL YSKQMHFNFV IYSAFGRSSP IDSSAALDEL 900 EGKCVGRQKK IVVAGKWCGK VWMSSQAHPL LLNKDSQEQE EESEFTAWIK PNLKSERLSQ 960 SSQAAGVASA ICKTSRKRKN NAENSSHVKE KSLEAGKMDE PSIGFPLSNC NKVIKRKRGS 1020 RRLKEENPES EDLDESSEEI QLSNSCNQIK KRHGTKLLKK DTFEADENLD ASSEDFPLSN 1080 SWKQIKSKRG ARKTNKETPE PLKSKKRSKQ QANSLVDDDE LEGGPSTRLR KRTTKACKAS 1140 GPRSTNAKPV LKKQQKDIKT KKVPPVKVPS KAKLKDEEAE YACDMEGCTM SFGSKQELAL 1200 HKKNICPVKG CGKKFFSHKY LVQHRRVHMD DRPLKCPWKG CKMTFKWAWA RTEHIRVHTG 1260 ARPYVCTEAG CGQTFRFVSD FSRHKRKTGH SPKKARG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 2e-75 | 1 | 358 | 12 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 2e-75 | 1 | 358 | 12 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1014 | 1023 | KRKRGSRRLK |
| 2 | 1049 | 1059 | KKRHGTKLLKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
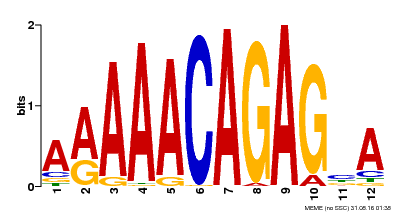
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_011072923.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011072922.1 | 0.0 | lysine-specific demethylase REF6 isoform X1 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A2G9I3H7 | 0.0 | A0A2G9I3H7_9LAMI; [histone H3]-lysine-36 demethylase | ||||
| STRING | Migut.I00266.1.p | 0.0 | (Erythranthe guttata) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



