 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.010G215700.1.p | ||||||||
| Common Name | Sb10g026200, SORBIDRAFT_10g026200 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 445aa MW: 47129.6 Da PI: 9.1554 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.1 | 8.1e-41 | 194 | 270 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cq+egC+adls ak+yhrrhkvCe h+ka+vv++ g++qrfCqqCsrfh lsefDe krsCr+rLa+hn+rrrk++
Sobic.010G215700.1.p 194 RCQAEGCKADLSGAKHYHRRHKVCEYHAKASVVAAGGKQQRFCQQCSRFHVLSEFDEVKRSCRKRLAEHNRRRRKPA 270
6**************************************************************************87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.4E-34 | 187 | 256 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.359 | 192 | 269 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.62E-38 | 193 | 272 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.0E-31 | 195 | 268 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 445 aa Download sequence Send to blast |
MMSGRLNVAT SASPGDDFPF ASMQQQQQPP PPPPPPPYVG FEHGVAGGGG GQRGAGGMGM 60 QQQQHLYDGL DFAAALQFQQ EAPHHHHHQL LTLPSSLGPM APPPLPMPMP MPLQMPMPMP 120 GMPGGDHVYQ ALGMVKREGG GGGGADAGRI GLNLGRRTYF SPGDMLAVDR LLMRSRLGGV 180 FGLGFGGAHH QPPRCQAEGC KADLSGAKHY HRRHKVCEYH AKASVVAAGG KQQRFCQQCS 240 RFHVLSEFDE VKRSCRKRLA EHNRRRRKPA TTASSKDAAS PPANKKPNGG SITSSYSTDN 300 KNLSTAKSTM SSNTSSVISC LDQAGNKQQQ LARPTLTLGA SQDNKDQHQH QQQLSTMLQV 360 QAAGGGHHQE QHFLTSLQVH NNGGCGGGNN NNILSCSSVC SSGALPSSAN GEVSDQNTTT 420 TTTNNGNGGS SNNNLHNLFE VDFM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-28 | 195 | 268 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 251 | 268 | KRSCRKRLAEHNRRRRKP |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in stems, leaf sheaths, and young panicles. {ECO:0000269|PubMed:16861571}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in stems, leaf sheaths, and young panicles. {ECO:0000269|PubMed:16861571}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
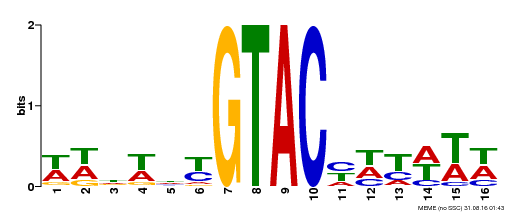
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.010G215700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT033937 | 0.0 | BT033937.1 Zea mays full-length cDNA clone ZM_BFc0033O21 mRNA, complete cds. | |||
| GenBank | BT034298 | 0.0 | BT034298.1 Zea mays full-length cDNA clone ZM_BFc0007A11 mRNA, complete cds. | |||
| GenBank | BT065712 | 0.0 | BT065712.1 Zea mays full-length cDNA clone ZM_BFc0041B17 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002437398.1 | 0.0 | squamosa promoter-binding-like protein 10 | ||||
| Swissprot | A2YFT9 | 1e-107 | SPL10_ORYSI; Squamosa promoter-binding-like protein 10 | ||||
| Swissprot | Q0DAE8 | 1e-107 | SPL10_ORYSJ; Squamosa promoter-binding-like protein 10 | ||||
| TrEMBL | C5Z790 | 0.0 | C5Z790_SORBI; Uncharacterized protein | ||||
| STRING | Sb10g026200.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2375 | 35 | 93 | Representative plant | OGRP97 | 17 | 230 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 3e-48 | squamosa promoter binding protein-like 8 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.010G215700.1.p |
| Entrez Gene | 8058456 |



