 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.008G014801.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 866aa MW: 97478.9 Da PI: 7.841 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.1 | 0.0012 | 557 | 579 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cg +F + +Lk+H++ H
Sobic.008G014801.1.p 557 HRCQECGSCFQKPAHLKQHMQGH 579
79******************987 PP
| |||||||
| 2 | zf-C2H2 | 22.2 | 3.6e-07 | 585 | 609 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s++rk++L+rH+ +H
Sobic.008G014801.1.p 585 FMCPieDCPFSYKRKDHLNRHMLKH 609
79*********************99 PP
| |||||||
| 3 | zf-C2H2 | 15.8 | 4.1e-05 | 614 | 639 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C+ C+++Fs k n++rH + H
Sobic.008G014801.1.p 614 FSCTvdGCDRRFSMKANMQRHVKEiH 639
899999****************9988 PP
| |||||||
| 4 | zf-C2H2 | 15.3 | 5.6e-05 | 651 | 675 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ C C+k+F+ +s Lk+H +H
Sobic.008G014801.1.p 651 FICReeGCNKVFRYSSKLKKHEESH 675
789999***************9988 PP
| |||||||
| 5 | zf-C2H2 | 13.2 | 0.00027 | 742 | 766 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C sFs+ksnL++H + H
Sobic.008G014801.1.p 742 KCTfeGCEHSFSNKSNLTKHVKAcH 766
799999***************9955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51375 | 5.305 | 35 | 65 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 2.5E-11 | 44 | 163 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 8.67 | 66 | 100 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.802 | 136 | 170 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 7.2E-5 | 141 | 164 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 3.2E-4 | 141 | 164 | IPR002885 | Pentatricopeptide repeat |
| SuperFamily | SSF48452 | 3.21E-5 | 147 | 174 | IPR011990 | Tetratricopeptide-like helical domain |
| Pfam | PF01535 | 0.15 | 169 | 193 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 10.03 | 198 | 232 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 3.2E-4 | 200 | 229 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 5.6E-5 | 200 | 233 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 2.5E-11 | 250 | 335 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 8.681 | 268 | 298 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 0.0012 | 273 | 298 | IPR002885 | Pentatricopeptide repeat |
| SuperFamily | SSF48452 | 3.21E-5 | 289 | 329 | IPR011990 | Tetratricopeptide-like helical domain |
| Pfam | PF13041 | 6.7E-12 | 298 | 346 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 12.617 | 299 | 333 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 1.0E-9 | 301 | 335 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 9.591 | 334 | 364 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 5.7E-4 | 336 | 363 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 6.38 | 370 | 400 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.017 | 373 | 397 | IPR002885 | Pentatricopeptide repeat |
| SuperFamily | SSF48452 | 3.21E-5 | 400 | 465 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 5.338 | 402 | 432 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 2.5E-11 | 403 | 466 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 7.311 | 436 | 470 | IPR002885 | Pentatricopeptide repeat |
| SMART | SM00355 | 22 | 511 | 533 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-4 | 556 | 578 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.1 | 557 | 579 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.258 | 557 | 584 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 559 | 581 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.54E-9 | 570 | 611 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 4.9E-10 | 579 | 611 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.182 | 585 | 614 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.6E-4 | 585 | 609 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 587 | 609 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.63E-8 | 595 | 644 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-8 | 612 | 641 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.759 | 614 | 644 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.0E-4 | 614 | 639 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 616 | 639 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-7 | 649 | 675 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.284 | 651 | 675 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0085 | 651 | 675 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 653 | 675 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.3 | 683 | 708 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 685 | 708 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 14 | 711 | 732 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 2.49E-8 | 726 | 783 | No hit | No description |
| SMART | SM00355 | 0.047 | 741 | 766 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.489 | 741 | 771 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 3.7E-6 | 742 | 766 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 743 | 766 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-5 | 767 | 794 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.515 | 772 | 798 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.91 | 772 | 798 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 774 | 798 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 866 aa Download sequence Send to blast |
MNRSVCHHLL AQCKKLRELQ RIHALAIAHG LHPHQQSVSC KIFRCYADFG RVADARKLFD 60 EIPNPDLISF TSLMSLHLQL DNQREAISLF ARVVAAGHRP DGFAVVGALS ASSGAGDQVV 120 GRAVHGLIFR LGLDGEVVVG NALIDMYSQC GKFESAVKVF DRMSLKDEVT WGSMLHGYIK 180 CAGVDSALSF FDQVPVRSVV AWTALITGHV QGRQPVRALE LFGRMVLEGH RPTHVTIVGV 240 LSACADIGAL DLGRVIHGYG SKCNASLNII VSNALMDMYA KSGHIEMAFS VFQEVQSKDS 300 FTWTTMISCC TVQGDGKKAL ELFQDMLRAG VVPNSVTFVS VLSACSHSGL IEEGIELFDR 360 MRQLYKIDPL LEHYGCMIDL LGRGGLLEEA EALIADMNVE PDIVIWRSLL SACLVRGNNR 420 LAEIAGKEIV KREPGDDGVY VLLWNMYASS NKWREAREMR QQMLSLKIFK KPGCSWIEID 480 GAVHEFLMCS VVDVDGDASV GQKGYRDIRR YKCEFCTVVR SKKCLIQAHM VAHHKDELDK 540 SEIYNSNGEK IVHEEEHRCQ ECGSCFQKPA HLKQHMQGHS HERLFMCPIE DCPFSYKRKD 600 HLNRHMLKHE GKLFSCTVDG CDRRFSMKAN MQRHVKEIHE DENASKSNQQ FICREEGCNK 660 VFRYSSKLKK HEESHVKLDY VEVLCGEPGC MKMFTNVEYL RAHSQSCHQY VQCEICGEKH 720 LKKNIKRHLQ SHDKAPSGER MKCTFEGCEH SFSNKSNLTK HVKACHDQLK PFKCQIAGCG 780 KAFTYKHVRD NHEKSGAHVY IEGDFEEIDE QLRARPRGGR KRKALTVETL TRKRVTIPGE 840 ASSLDDGEEY LRWLLSGGDS SRETQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5iww_D | 6e-54 | 9 | 413 | 15 | 331 | PLS9-PPR |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 814 | 822 | RPRGGRKRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
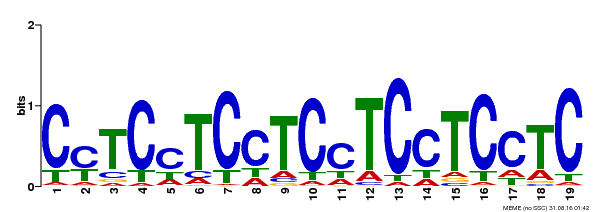
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.008G014801.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FN431668 | 0.0 | FN431668.1 Saccharum hybrid cultivar BAC clone Sh253G12, cultivar R570. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002441722.2 | 0.0 | pentatricopeptide repeat-containing protein At2g22410, mitochondrial isoform X1 | ||||
| TrEMBL | A0A1Z5R4A5 | 0.0 | A0A1Z5R4A5_SORBI; Uncharacterized protein | ||||
| STRING | Si020193m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 | Representative plant | OGRP2981 | 16 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-102 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.008G014801.1.p |



