 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G72050.2 | ||||||||
| Common Name | F28P5.6, TFIIIA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 412aa MW: 46652.9 Da PI: 7.49 | ||||||||
| Description | transcription factor IIIA | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.2 | 0.0012 | 20 | 42 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C++Cg s s +++ ++Hi++H
AT1G72050.2 20 YLCQYCGISRSKNYLITKHIQSH 42
56********************6 PP
| |||||||
| 2 | zf-C2H2 | 20.1 | 1.7e-06 | 66 | 88 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cg F+ + +Lk+H+++H
AT1G72050.2 66 HTCQECGAEFKKPAHLKQHMQSH 88
79*******************99 PP
| |||||||
| 3 | zf-C2H2 | 15.4 | 5.5e-05 | 94 | 118 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C dC s++rk++L+rH+ tH
AT1G72050.2 94 FTCYvdDCAASYRRKDHLNRHLLTH 118
789999*****************99 PP
| |||||||
| 4 | zf-C2H2 | 21.9 | 4.6e-07 | 216 | 239 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+C+ Cgk F+ +s+L++H+ +H
AT1G72050.2 216 VCKeiGCGKAFKYPSQLQKHQDSH 239
69999****************998 PP
| |||||||
| 5 | zf-C2H2 | 18.9 | 4.2e-06 | 306 | 330 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C+ +Fs snL++H++ H
AT1G72050.2 306 KCEveGCSSTFSKASNLQKHMKAvH 330
79999****************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 22 | 20 | 42 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 8.2E-5 | 65 | 85 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 12.819 | 66 | 93 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0015 | 66 | 88 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 68 | 88 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.41E-9 | 79 | 120 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.7E-11 | 86 | 120 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.91 | 94 | 123 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.11 | 94 | 118 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 96 | 118 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.89 | 123 | 148 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.723 | 123 | 153 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 125 | 148 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.7E-5 | 210 | 241 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-6 | 210 | 239 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0063 | 215 | 239 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.533 | 215 | 244 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 217 | 239 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.95 | 247 | 272 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 249 | 272 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 22 | 275 | 296 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 3.35E-8 | 286 | 340 | No hit | No description |
| PROSITE profile | PS50157 | 12.258 | 305 | 335 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5.3E-4 | 305 | 330 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.5E-7 | 306 | 330 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 307 | 330 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 7.5E-5 | 331 | 357 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 5.7 | 336 | 362 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 338 | 362 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007275 | Biological Process | multicellular organism development | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000005 | anatomy | cultured plant cell | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020094 | anatomy | plant egg cell | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 412 aa Download sequence Send to blast |
MAEEAKVDVK TSAKKDIRNY LCQYCGISRS KNYLITKHIQ SHHQMELEEE RDDEACEVDE 60 ESSSNHTCQE CGAEFKKPAH LKQHMQSHSL ERSFTCYVDD CAASYRRKDH LNRHLLTHKG 120 KLFKCPKENC KSEFSVQGNV GRHVKKYHSN DNRDKDNTGL GDGDKDNTCK GDDDKEKSGS 180 GGCEKENEGN GGSGKDNNGN GDSQPAECST GQKQVVCKEI GCGKAFKYPS QLQKHQDSHV 240 KLDSVEAFCS EPGCMKYFTN EECLKSHIRS CHQHINCEIC GSKHLKKNIK RHLRTHDEDS 300 SPGEIKCEVE GCSSTFSKAS NLQKHMKAVH DDIRPFVCGF PGCGMRFAYK HVRNKHENSG 360 YHVYTCGDFV ETDEDFTSRP RGGLKRKQVT AEMLVRKRVM PPRFDAEEHE TC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wjq_D | 3e-13 | 18 | 365 | 7 | 284 | Zinc finger protein 568 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.66854 | 0.0 | flower | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 145337437 | 0.0 | ||||
| Genevisible | 256344_at | 0.0 | ||||
| Expression Atlas | AT1G72050 | - | ||||
| AtGenExpress | AT1G72050 | - | ||||
| ATTED-II | AT1G72050 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Both isoforms 1 and 2 transcripts are expressed in seedlings and both TFIIIA and TFIIIA-C are present at similar ratios in 10-day-old seedlings (at protein level). In correlation with the reorganization of 5S rDNA chromatin to a mature state, full-length isoform 1 protein TFIIIA with transcriptional activity accumulates and permits de novo transcription of 5S rRNA. Isoforms 1 and 2 transcripts accumulate in various tissues of the reproductive phase, including flowers and siliques, but only isoform 2 is present in mature seeds. In flowers, both TFIIIA and TFIIIA-C are present at similar ratios (at protein level). Very low amounts of TFIIIA are found in extracts from fresh siliques compared to TFIIIA-C. The amount of full-length TFIIIA protein progressively increases from fresh siliques to seeds concomitant with lower proportions of the shorter TFIIIA-C protein (at protein level) thus leading to 5S rRNA accumulation in the seed. {ECO:0000269|PubMed:22353599}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in seedlings, flowers, siliques and seeds. {ECO:0000269|PubMed:22353599}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a transcriptional factor TFIIIA required for transcription of 5S rRNA gene. 5S rRNA is the smallest constituent of the ribosome. Work on one of the gene models AT1G72050.2 showed that it encodes a protein with nine Cys(2)-His(2)-type zinc fingers, a characteristic feature of TFIIIA proteins. AT1G72050.2 also contains a 23 amino acid spacer between fingers 1 and 2, a 66 amino acid spacer between fingers 4 and 5, and a 50 amino acid non-finger C-terminal tail. in vitro assay demonstrated that AT1g72050.2 binds to 5S rDNA and efficiently stimulates the transcription of 5S rRNA. AT1g72050.2 also binds to 5S rRNA in vitro. AT1g72050.2 is located at several nuclear foci including the nucleolus and is absent from the cytoplasm. | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | 27203113 | Download |
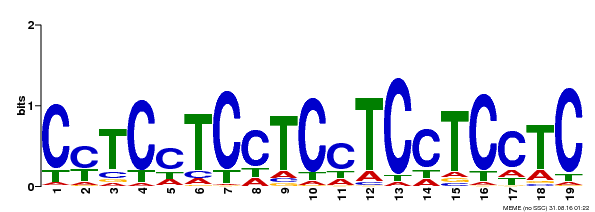
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G72050.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| IntAct | Search Q84MZ4 | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Lethal. {ECO:0000269|PubMed:22353599}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G72050 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ630478 | 0.0 | AJ630478.1 Arabidopsis thaliana mRNA for hypothetical protein, clone At1g72050D. | |||
| GenBank | AY186610 | 0.0 | AY186610.1 Arabidopsis thaliana transcription factor IIIA (TFIIIA) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001322779.1 | 0.0 | transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 0.0 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A1P8AV27 | 0.0 | A0A1P8AV27_ARATH; Transcription factor IIIA | ||||
| STRING | AT1G72050.2 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4714 | 28 | 52 | Representative plant | OGRP2981 | 16 | 27 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G72050.2 |
| Entrez Gene | 843536 |
| iHOP | AT1G72050 |
| wikigenes | AT1G72050 |



