 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.004G283201.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 334aa MW: 36064.4 Da PI: 9.5885 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 57.7 | 3e-18 | 163 | 213 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ ykGVr++ grWv+e+r+p g ++r++lg+f tae+Aa+a++ a+++l+g
Sobic.004G283201.1.p 163 PVYKGVRRRN-PGRWVCEVREP--HG-KQRIWLGTFETAEMAARAHDVAALALRG 213
68*****887.9******9996..33.5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 1.6E-13 | 163 | 213 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.089 | 164 | 221 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.9E-21 | 164 | 222 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 6.0E-28 | 164 | 227 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.0E-30 | 164 | 223 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.61E-20 | 165 | 223 | No hit | No description |
| PRINTS | PR00367 | 5.0E-8 | 165 | 176 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.0E-8 | 187 | 203 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009631 | Biological Process | cold acclimation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MPARPHPRPP APRAPAGSDP LRARRARPLS PSPEHPLPDR PIKSHYLAGA SRAAPRWNSA 60 RGHVALHALT YITRHLSLQI SNQMLRHTFL LPPARHGHHH TRPSPMDAAS FSDYSSGTPS 120 PVGGVSGGVD GDDGGSSSSY MTVSSAPPKR RAGRTKFKET RHPVYKGVRR RNPGRWVCEV 180 REPHGKQRIW LGTFETAEMA ARAHDVAALA LRGRAACLNF ADSPRLLRVP PMGSGHDEIR 240 RAAAVAADQF RPAPDRQGNV ATAEEAADTT PLDATTQSVV DDPYCIIDDR LDFGMQGYLD 300 MAQGMLIDPP PMAGSSTSGG DDDDDGEVKL WSY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 5e-14 | 162 | 220 | 12 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity- and dehydration-inducible transcription (By similarity). {ECO:0000250}. | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity- and dehydration-inducible transcription (By similarity). {ECO:0000250}. | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity- and dehydration-inducible transcription (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00644 | PBM | Transfer from LOC_Os02g45450 | Download |
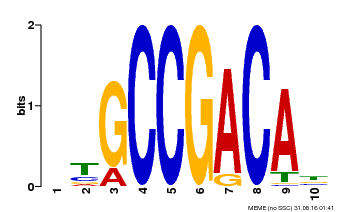
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.004G283201.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ727873 | 0.0 | KJ727873.1 Zea mays clone pUT5966 AP2-EREBP transcription factor (EREB16) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002454484.2 | 0.0 | dehydration-responsive element-binding protein 1G | ||||
| Swissprot | A2XWL6 | 6e-96 | DRE1E_ORYSI; Dehydration-responsive element-binding protein 1E | ||||
| Swissprot | Q6EP77 | 8e-96 | DRE1G_ORYSJ; Dehydration-responsive element-binding protein 1G | ||||
| Swissprot | Q8H273 | 6e-96 | DRE1E_ORYSJ; Dehydration-responsive element-binding protein 1E | ||||
| TrEMBL | A0A1Z5RPX1 | 0.0 | A0A1Z5RPX1_SORBI; Uncharacterized protein | ||||
| STRING | Sb04g031950.1 | 1e-164 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP196 | 36 | 261 | Representative plant | OGRP6 | 16 | 1718 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G25470.1 | 7e-36 | C-repeat/DRE binding factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.004G283201.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



