 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.003G394900.1.p | ||||||||
| Common Name | Sb03g043210, SORBIDRAFT_03g043210 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1346aa MW: 149029 Da PI: 8.7311 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 14 | 0.00015 | 1307 | 1333 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+Cp Cg++F+ s++ rH r+ H
Sobic.003G394900.1.p 1307 YVCPepECGQTFRFVSDFSRHKRRtgH 1333
99********************98666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 7.7E-15 | 18 | 59 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.481 | 19 | 60 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 9.9E-13 | 20 | 53 | IPR003349 | JmjN domain |
| SMART | SM00558 | 2.3E-48 | 184 | 353 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 35.433 | 187 | 353 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.35E-25 | 200 | 350 | No hit | No description |
| Pfam | PF02373 | 2.2E-38 | 217 | 336 | IPR003347 | JmjC domain |
| SMART | SM00355 | 34 | 1224 | 1246 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.094 | 1247 | 1276 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 13 | 1247 | 1271 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1249 | 1271 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.68E-15 | 1258 | 1318 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-9 | 1275 | 1301 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.635 | 1277 | 1306 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 7.8E-4 | 1277 | 1301 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1279 | 1301 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-10 | 1302 | 1329 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.74 | 1307 | 1333 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.323 | 1307 | 1338 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1309 | 1333 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1346 aa Download sequence Send to blast |
MASSQGEPVP PWLKSLPLAP EFRPTAAEFV DPIAYLLKIE PAAAPFGICK IVPPLPPPPK 60 RATLGNLSRS FAALHPDDPT PTFPTRHQQL GLCPRRPRPA LKPVWLSSHR YTLPKFEAKA 120 GASRKALLAR LSVPATKQLS PLDVEALFWR SSADRPVVVE YASDMPGSGF APCAARPTQL 180 PAANVGETAW NMRGVARSPA SLLRFVREEV PGVTSPMLYV GMMFSWFAWH VEDHDLHSLN 240 YMHYGAPKTW YAVPRDAALA FEEVVRVHGY GGEVNSLETF AMLGDKTTVM SPQVLVDSGI 300 PCCRLVQNAG EFVVTFPRAY HSGFSHGFNC GEASNIATPE WLRVAKEAAV RRASINRPPM 360 VSHYQLLYEL ALSLCLRDPS NGTMEPRSCR LKEKKKSEGD QLIKKIFVQN VIEDNKLLGH 420 FLSDGSSCII LPVNYSDGSP LSTLLSKFQS TTDSRISHDQ CSKTEAPKDS RCLPKDQADK 480 NWELSSSNKI SLSVCSGKTV PPTTCIYDCA NMSASSYTHN TENNKGGMNS AAGLLDQGLL 540 SCVACGILSF SCVAVIKPRE CAAKWLMTAD SSLINDRLAS SGEHHMIDGL QGGRTTGGIL 600 RSDSEMNGNS IISDADDVPL NGCSALDLLA SAYGDPSDSD EDVMNKKIQI PNVSNELINH 660 TVESQPNTSN NGDCDGTKVS SSSKESQQGP TSQNSKCIGN SNTLNGPKGV RTRNKDLLKM 720 VLSEGFQPKD IYSETHKKVQ CEPSSSNKTS TESPCSTEYH VSHNSATICM DNNRGSMTMV 780 NNLVTSVVKP DKDSSRMHVF CLEHAIEVEK QLQAIGGADI FLLCRPEYPR IEVEARLLAE 840 EMEFVYDWKD ILFKEATIED REKIQEVVQD EEAIPTNSDW AVKLGINLYY SANLAKSPLY 900 NKQVPYNRVI YEAFGYGSPN DSPVKLKTYS RRQGRAKKIV LAGRWCGKVW MSNQVHPYLA 960 DRIKIHEPEE TDETFPSDQK SNAEPVEDSS RQAASTRKSS SRAIEGKISK REKESLEKAN 1020 AKKPKITEED NSKSLEGTAE ASTQSRTALE KTSRKEKEHV EKANTKKLKH TEKVSEAIKG 1080 PSEASYPAPA GMVVRSSSRI ANRKSMLKSR MDEEDIGSTN RPKSKVEDDK DNPAGRSTAK 1140 SLRQKTKLDA KKKTKETRVE KRKAPSPASQ KDEEEQSYEG CSITKQRLSL RKKGAKIEEK 1200 QQQMEKSRYR GRAPPSSPKS KEEYACDVEG CSMSFGTEEA LSLHKNDICP EKGCCRKFFS 1260 HKYLLQHRKV HTDDRPLKCP WKGCSMAFKW PWARTEHMRV HTGDRPYVCP EPECGQTFRF 1320 VSDFSRHKRR TGHAGMPAKK AKAKK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 2e-76 | 12 | 374 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 2e-76 | 12 | 374 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaves and flag leaves. Expressed at low levels in roots, shoots, stems and panicles. {ECO:0000269|PubMed:24280387}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3 with a specific activity for H3K27me3 and H3K27me2. No activity on H3K4me3, H3K9me3, H3K27me1 and H3K36me3. Involved in biotic stress response. May demethylate H3K27me3-marked defense-related genes and increase their basal and induced expression levels during pathogen infection. {ECO:0000269|PubMed:24280387}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
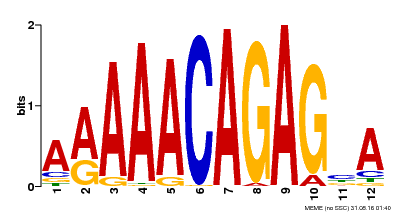
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.003G394900.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt stress, abscisic acid (ABA), jasmonic acid (JA), the ethylene precursor ACC and infection by the bacterial pathogen Xanthomonas oryzae pv. oryzae. {ECO:0000269|PubMed:24280387}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU949054 | 0.0 | EU949054.1 Zea mays clone 394823 mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021310712.1 | 0.0 | lysine-specific demethylase JMJ705 | ||||
| Swissprot | Q5N712 | 0.0 | JM705_ORYSJ; Lysine-specific demethylase JMJ705 | ||||
| TrEMBL | A0A1B6Q7Q4 | 0.0 | A0A1B6Q7Q4_SORBI; Uncharacterized protein | ||||
| STRING | Sb03g043210.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5346 | 32 | 46 | Representative plant | OGRP4401 | 12 | 17 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.003G394900.1.p |
| Entrez Gene | 8056139 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



