 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.003G383700.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | LBD | ||||||||
| Protein Properties | Length: 333aa MW: 34635.3 Da PI: 9.0403 | ||||||||
| Description | LBD family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF260 | 137.3 | 5.3e-43 | 106 | 204 | 1 | 99 |
DUF260 1 aCaaCkvlrrkCakdCvlapyfpaeqpkkfanvhklFGasnvlkllkalpeeeredamsslvyeAearardPvyGavgvilklqqqleq 89
+CaaCk+lrrkC++dCv+apyfp ++p+kf vh++FGasnv+kll++l++ +reda++sl+yeA++r+rdPvyG+vgvi+ lq++l+q
Sobic.003G383700.2.p 106 PCAACKFLRRKCQPDCVFAPYFPPDNPQKFVHVHRVFGASNVTKLLNELHPFQREDAVNSLAYEADMRLRDPVYGCVGVISILQHNLRQ 194
7**************************************************************************************** PP
DUF260 90 lkaelallke 99
l+++la++k
Sobic.003G383700.2.p 195 LQQDLARAKY 204
*****99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50891 | 26.729 | 105 | 206 | IPR004883 | Lateral organ boundaries, LOB |
| Pfam | PF03195 | 6.8E-43 | 106 | 202 | IPR004883 | Lateral organ boundaries, LOB |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009799 | Biological Process | specification of symmetry | ||||
| GO:0009944 | Biological Process | polarity specification of adaxial/abaxial axis | ||||
| GO:0009954 | Biological Process | proximal/distal pattern formation | ||||
| GO:0048441 | Biological Process | petal development | ||||
| GO:0005654 | Cellular Component | nucleoplasm | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 333 aa Download sequence Send to blast |
MVLHKAPVSK GLFFFLCPPR LRALSLRASQ LGDDYAKCPG LLQARSLLLY TSFPGLLRSS 60 DSSSPATSTA SRRMASSVPA PSGSVITVAS SSSSAAAAAV CGTGSPCAAC KFLRRKCQPD 120 CVFAPYFPPD NPQKFVHVHR VFGASNVTKL LNELHPFQRE DAVNSLAYEA DMRLRDPVYG 180 CVGVISILQH NLRQLQQDLA RAKYELSKYQ AAAAASASTA APNGPQAMAE FIGNAMPNGA 240 HNFINIGHSA ALGSLGGSAT VFGQEQFANA QMLSRSYDGE PIARLGINGG YEFGYSTAMG 300 GSGAVSGLGT LGISPFLKSG TAGGDEKPSG GY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5ly0_A | 6e-53 | 96 | 222 | 1 | 127 | LOB family transfactor Ramosa2.1 |
| 5ly0_B | 6e-53 | 96 | 222 | 1 | 127 | LOB family transfactor Ramosa2.1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.5472 | 0.0 | pollen | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: In embryo sac, detected as early as the one-nucleus stage. In older embryo sacs, highest expression in antipodal cells. {ECO:0000269|PubMed:17209126}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaves, leaf primordia, immature ears, immature tassels, whole ovules, silk and husk leaves. Found on the adaxial side of organs. {ECO:0000269|PubMed:17209126}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Promotes the switch from proliferation to differentiation in the embryo sac. Negative regulator of cell proliferation in the adaxial side of leaves. Regulates the formation of a symmetric lamina and the establishment of venation. Interacts directly with RS2 (rough sheath 2) to repress some knox homeobox genes. {ECO:0000269|PubMed:17209126}. | |||||
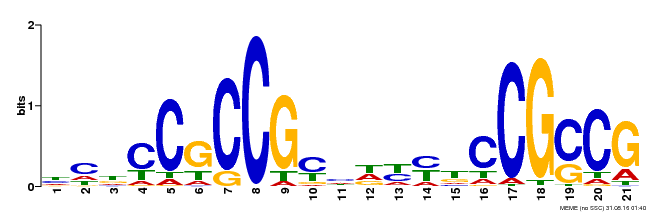
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00211 | DAP | Transfer from AT1G65620 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.003G383700.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY940681 | 0.0 | AY940681.1 Zea mays ASYMMETRIC LEAVES2 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021311343.1 | 0.0 | LOB domain-containing protein 6 isoform X2 | ||||
| Swissprot | Q32SG3 | 1e-166 | LBD6_MAIZE; LOB domain-containing protein 6 | ||||
| TrEMBL | A0A1W0W0W9 | 0.0 | A0A1W0W0W9_SORBI; Uncharacterized protein | ||||
| STRING | Sb03g042310.2 | 0.0 | (Sorghum bicolor) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G65620.4 | 1e-57 | LBD family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.003G383700.2.p |



