 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.001G287600.1.p | ||||||||
| Common Name | Sb01g028230, SORBIDRAFT_01g028230 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 708aa MW: 75519.3 Da PI: 6.5593 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39 | 1.4e-12 | 529 | 574 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ K++Ka+ L A+ YI++L
Sobic.001G287600.1.p 529 NHVEAERQRREKLNQRFYALRAVVPNV-----SKMDKASLLGDAISYINEL 574
799***********************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 8.3E-53 | 62 | 242 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.39 | 525 | 574 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 2.09E-18 | 528 | 596 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.92E-14 | 528 | 578 | No hit | No description |
| Pfam | PF00010 | 4.7E-10 | 529 | 574 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 9.5E-19 | 529 | 596 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 3.6E-16 | 531 | 580 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009269 | Biological Process | response to desiccation | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009867 | Biological Process | jasmonic acid mediated signaling pathway | ||||
| GO:0009963 | Biological Process | positive regulation of flavonoid biosynthetic process | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0043619 | Biological Process | regulation of transcription from RNA polymerase II promoter in response to oxidative stress | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051090 | Biological Process | regulation of sequence-specific DNA binding transcription factor activity | ||||
| GO:2000068 | Biological Process | regulation of defense response to insect | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 708 aa Download sequence Send to blast |
MNLWTDDNAS MMEAFMASAD LPTFPWGATA GGGNSSAAAA TPPPPPQMPA AAMAPGFNQD 60 TLQQRLQAMI EGSSETWTYA IFWQSSLDAA TGASLLGWGD GYYKGCDDDK RKQRPLTPAA 120 QAEQEHRKRV LRELNSLISG AAAAPDEAVE EEVTDTEWFF LVSMTQSFLN GSGLPGQALF 180 AGQPTWIASG LSSAPCERAR QAYNFGLRTM VCFPVGTGVL ELGSTDVVFQ TAESMAKIRS 240 LFGGGAGGGS WPPPQAPSHQ QPAAGPDQAE TDLWLADAPV MDIKDSMSHP SAEISVSKPP 300 PPPPPPQIHF ENASTSTLTE NPSPSVHAAP PQPAPAAAPQ RQHQHQNQAH QGPFRRELNF 360 SDFASTNPSS LAATPPFFKP ESGEILSFGA DSNARRNPSP APPAATASLT TAPGSLFSQH 420 TATMTQAAAA NDAKNNNKRS MEATSRASNT NHHPAATANE GMLSFSSAPT TRPSTGTGAP 480 AKSESDHSDL DASVREVESS RVVAPPPEAE KRPRKRGRKP ANGREEPLNH VEAERQRREK 540 LNQRFYALRA VVPNVSKMDK ASLLGDAISY INELRGKLTS LESDKDTLQA QIEALKKERD 600 ARPPAHAAGL GGHDGGPRCH AVEIDAKILG LEAMIRVQCH KRNHPSARLM TALRELDLDV 660 YHASVSVVKD LMIQQVAVKM ASRIYSQDQL NAALYSRLAE PGSAMGR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 3e-63 | 55 | 242 | 2 | 192 | Transcription factor MYC3 |
| 4rqw_B | 3e-63 | 55 | 242 | 2 | 192 | Transcription factor MYC3 |
| 4rs9_A | 3e-63 | 55 | 242 | 2 | 192 | Transcription factor MYC3 |
| 4yz6_A | 3e-63 | 55 | 242 | 2 | 192 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 510 | 518 | KRPRKRGRK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.15724 | 0.0 | leaf| ovary | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in spikelets and floral organs. {ECO:0000269|PubMed:24647160}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in jasmonate (JA) signaling pathway during spikelet development. Binds to the G2 region G-box (5'-CACGTG-3') of the MADS1 promoter and thus directly regulates the expression of MADS1. Its function in MADS1 activation is abolished by TIFY3/JAZ1 which directly target MYC2 during spikelet development. {ECO:0000269|PubMed:24647160}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00084 | PBM | Transfer from AT1G32640 | Download |
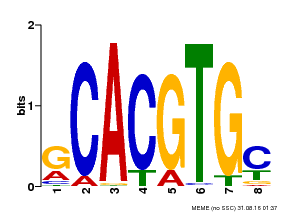
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.001G287600.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF061107 | 0.0 | AF061107.1 Zea mays transcription factor MYC7E mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002467448.1 | 0.0 | transcription factor MYC2 | ||||
| Swissprot | Q336P5 | 0.0 | MYC2_ORYSJ; Transcription factor MYC2 | ||||
| TrEMBL | A0A1B6QMQ5 | 0.0 | A0A1B6QMQ5_SORBI; Uncharacterized protein | ||||
| STRING | Sb01g028230.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4452 | 31 | 57 | Representative plant | OGRP3733 | 12 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G17880.1 | 5e-67 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.001G287600.1.p |
| Entrez Gene | 8055143 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



