 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_05436.1_g00001.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 391aa MW: 44740.4 Da PI: 5.0476 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 109.6 | 2.3e-34 | 17 | 108 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFk 87
Fl+k+ye+++d++ ++++sws++++sf+v+++ ef++++Lp++Fkh+nf+SF+RQLn+YgF+k + e+ weF+++ F
Rsa1.0_05436.1_g00001.1 17 FLTKTYEMVDDSSSDSIVSWSQSNRSFIVWNPPEFSRDLLPRFFKHNNFSSFIRQLNTYGFRKADPEK---------WEFANDDFV 93
9**************************************************************99988.........********* PP
XXXXXXXXXXXXXXX CS
HSF_DNA-bind 88 kgkkellekikrkks 102
+g+ +l+++i+r+k
Rsa1.0_05436.1_g00001.1 94 RGQPHLMKNIHRRKP 108
************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 1.1E-37 | 9 | 101 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 4.35E-35 | 12 | 106 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 6.8E-62 | 13 | 106 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 9.3E-31 | 17 | 106 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 5.0E-19 | 17 | 40 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 5.0E-19 | 55 | 67 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 5.0E-19 | 68 | 80 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 391 aa Download sequence Send to blast |
MDESNHGGSS SSSLPPFLTK TYEMVDDSSS DSIVSWSQSN RSFIVWNPPE FSRDLLPRFF 60 KHNNFSSFIR QLNTYGFRKA DPEKWEFAND DFVRGQPHLM KNIHRRKPVH SHSQPNLQAQ 120 QNPLTDSERQ RMSSQIERLT KEKEGLLQKL HKQEQERDQF EQQVKKLKDQ LQHMEKRQKT 180 MDSYVSQVVG TPELALNLSP CLPETNERKR RLPRIGFEQD NQTCVVVREE GSTSHSDETE 240 HQVEELESSI AVWESLVSDD SCEGMAQSTR SLMTLDVDDS STCPESPPLS CVQLSIDTCP 300 KSPPSQKIVD MNSEPDHVSK EQNTAVPTTA PAAGVNDVFW QQLLTENPDS AKQREVQPER 360 KEDKGEDQSE KCWWNSRNVN AITEQLGHLT S |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 7e-24 | 7 | 106 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 7e-24 | 7 | 106 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 7e-24 | 7 | 106 | 19 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 206 | 211 | ERKRRL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds DNA sequence 5'-AGAAnnTTCT-3' known as heat shock promoter elements (HSE). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
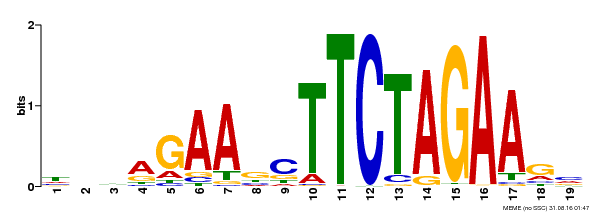
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_05436.1_g00001.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018454586.1 | 0.0 | PREDICTED: heat stress transcription factor A-4a-like | ||||
| Swissprot | O49403 | 0.0 | HFA4A_ARATH; Heat stress transcription factor A-4a | ||||
| TrEMBL | A0A398ARZ9 | 0.0 | A0A398ARZ9_BRACM; Uncharacterized protein | ||||
| STRING | Bo1g020010.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2853 | 28 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 0.0 | heat shock transcription factor A4A | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



