 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_02153.1_g00007.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 333aa MW: 38024.8 Da PI: 6.7137 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 165.3 | 2.1e-51 | 11 | 139 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk..kvka.eekewyfFskrdkkyatgkrknratksgyW 83
+ppGfrFhPt+eel+ +yL+kk+++ k++l +vi++vd++k+ePwd+++ k+ + +++wyfFs++dkky+tg+r+nrat+ g+W
Rsa1.0_02153.1_g00007.1 11 VPPGFRFHPTEEELLEYYLRKKISNIKIDL-DVIRDVDLNKLEPWDIQEmcKIGTtPQNDWYFFSHKDKKYPTGTRTNRATTVGFW 95
69****************************.9***************962433332455*************************** PP
NAM 84 katgkdkevlskkgelvglkktLvfykgrapkgektdWvmheyrl 128
katg+dk ++s +g +g++ktLvfykgrap+g+k+dW+mheyrl
Rsa1.0_02153.1_g00007.1 96 KATGRDKIIYS-NGDRIGMRKTLVFYKGRAPHGQKSDWIMHEYRL 139
***********.9999***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 6.93E-57 | 8 | 176 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 55.03 | 11 | 176 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.2E-27 | 12 | 139 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009834 | Biological Process | plant-type secondary cell wall biogenesis | ||||
| GO:0009901 | Biological Process | anther dehiscence | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 333 aa Download sequence Send to blast |
MKISVNGQSK VPPGFRFHPT EEELLEYYLR KKISNIKIDL DVIRDVDLNK LEPWDIQEMC 60 KIGTTPQNDW YFFSHKDKKY PTGTRTNRAT TVGFWKATGR DKIIYSNGDR IGMRKTLVFY 120 KGRAPHGQKS DWIMHEYRLD ENVLASSSNH DDDVSVETCE VIGGDEGWVV CRVFMKKSLC 180 KTVTMSSPPR SVKTSPETIE ELFEVMGQSC KEEITVDPFL KLPNNTIASC QRLMDDQVHN 240 YQVPSFGTSW ATLDRLVASQ LNGPNSYSIP SVDDIPQSPL HGLNRPGFYN TGLTQHYFTT 300 EMDIWNDTDF GRTTTSSSNP YGAMYQTVVN NRK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 7e-48 | 8 | 178 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 7e-48 | 8 | 178 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 7e-48 | 8 | 178 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 7e-48 | 8 | 178 | 14 | 167 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 9e-48 | 8 | 178 | 17 | 170 | NAC domain-containing protein 19 |
| 3swm_B | 9e-48 | 8 | 178 | 17 | 170 | NAC domain-containing protein 19 |
| 3swm_C | 9e-48 | 8 | 178 | 17 | 170 | NAC domain-containing protein 19 |
| 3swm_D | 9e-48 | 8 | 178 | 17 | 170 | NAC domain-containing protein 19 |
| 3swp_A | 9e-48 | 8 | 178 | 17 | 170 | NAC domain-containing protein 19 |
| 3swp_B | 9e-48 | 8 | 178 | 17 | 170 | NAC domain-containing protein 19 |
| 3swp_C | 9e-48 | 8 | 178 | 17 | 170 | NAC domain-containing protein 19 |
| 3swp_D | 9e-48 | 8 | 178 | 17 | 170 | NAC domain-containing protein 19 |
| 4dul_A | 7e-48 | 8 | 178 | 14 | 167 | NAC domain-containing protein 19 |
| 4dul_B | 7e-48 | 8 | 178 | 14 | 167 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator of genes involved in biosynthesis of secondary walls. Together with NST1, required for the secondary cell wall thickening of the anther endocethium, which is necessary for anther dehiscence. May also regulate the secondary cell wall lignification of other tissues such as tracheary elements. {ECO:0000269|PubMed:16214898}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
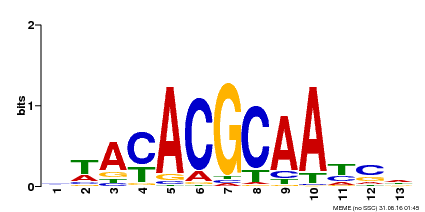
| Motif ID | Method | Source | Motif file |
| MP00015 | PBM | Transfer from AT3G61910 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_02153.1_g00007.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018465538.1 | 0.0 | PREDICTED: NAC domain-containing protein 66-like | ||||
| Swissprot | Q9M274 | 1e-180 | NAC66_ARATH; NAC domain-containing protein 66 | ||||
| TrEMBL | A0A3P6BF00 | 0.0 | A0A3P6BF00_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4CGZ5 | 0.0 | M4CGZ5_BRARP; Uncharacterized protein | ||||
| STRING | Bra003478.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM778 | 28 | 124 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G61910.1 | 1e-174 | NAC domain protein 66 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



