 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_01910.1_g00002.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 684aa MW: 75278.8 Da PI: 6.1858 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 51.3 | 2.3e-16 | 395 | 433 | 2 | 40 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
++k+CnCkkskClk+YCeCfaag++C e C+C dC+N+
Rsa1.0_01910.1_g00002.1 395 SCKRCNCKKSKCLKLYCECFAAGVYCIEPCSCVDCFNRP 433
689**********************************75 PP
| |||||||
| 2 | TCR | 52.8 | 8e-17 | 480 | 519 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
++k+gCnCkks+ClkkYCeCf++g+ Cs +C+Ce+CkN
Rsa1.0_01910.1_g00002.1 480 RHKRGCNCKKSNCLKKYCECFQSGVGCSINCRCEGCKNAF 519
589***********************************75 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 5.4E-19 | 394 | 435 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 37.515 | 395 | 520 | IPR005172 | CRC domain |
| Pfam | PF03638 | 5.7E-12 | 397 | 432 | IPR005172 | CRC domain |
| SMART | SM01114 | 5.7E-17 | 480 | 521 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 2.5E-12 | 482 | 518 | IPR005172 | CRC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009934 | Biological Process | regulation of meristem structural organization | ||||
| GO:0048444 | Biological Process | floral organ morphogenesis | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 684 aa Download sequence Send to blast |
MDSSQKKKKI ETPTPKSKFE DSPVFNFINN LSPIETVRSI PTVQTFNSLS FTSPPPVFTS 60 PHASFHRESR FFRCHNSVER SKALESLDST QEVVGGASGE VDLNKDATLE EEEEEEEEET 120 SCGHVDSPPG PGDIVTQVLL DPSGGAPRGE DNASSSSEDV RKILDAQEEN DTPGCGRLMA 180 DAAELLVFRS PNDSEAFGCL VDKISSSERR FCAGVKLTKQ HDITKDVPAN DNEPLALVPN 240 QSSVLSLHRG SMRRRCLDFE VLGKRKKDIA DDDKQTLGDN KAESSSKCVV PGIGLHLNAI 300 PMSSKAIKIN TRHEGSASVE IQKSFPGSIT PVHSQDIMQE TLVQAETEAG EGLAVEEETP 360 KPLVFEELNQ DSLQKKKQVF FLEKRKVEEP GEGDSCKRCN CKKSKCLKLY CECFAAGVYC 420 IEPCSCVDCF NRPIHEDTVL ATRKQIESRN PLAFAPKVIR NSDSIMETSD DASKTPASAR 480 HKRGCNCKKS NCLKKYCECF QSGVGCSINC RCEGCKNAFG KKEAYLLAIM ESKQEEDVSK 540 IQQNSDLPKE VEQNPSSDQP STPLPPYRHM VVHQPFLSKN RLPPTQFFLG AGSSSFRKPD 600 RDLTQARIDK KPLETVAEDK TEIMPEILSD TPITTVKAIS PNSKRVSPPH IGSSESGSIL 660 GKRTNGRKLI LRSIPAFPSL NPNQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 2e-16 | 399 | 517 | 14 | 120 | Protein lin-54 homolog |
| 5fd3_B | 2e-16 | 399 | 517 | 14 | 120 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 252 | 266 | RRRCLDFEVLGKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable floral-specific cell division component, required for proper organ formation in flowers. Regulates the floral meristem cell division and the inflorescence meristem organization. Plays a role in development of both male and female reproductive tissues. {ECO:0000269|PubMed:10769244, ECO:0000269|PubMed:10769245, ECO:0000269|PubMed:9043081, ECO:0000269|PubMed:9725857}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00370 | DAP | Transfer from AT3G22780 | Download |
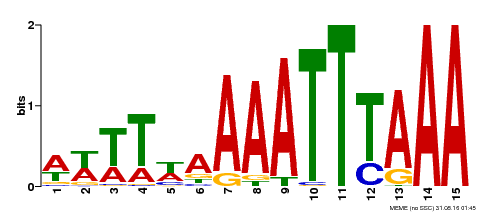
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_01910.1_g00002.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018493471.1 | 0.0 | PREDICTED: CRC domain-containing protein TSO1 | ||||
| Swissprot | Q9LUI3 | 0.0 | TSO1_ARATH; CRC domain-containing protein TSO1 | ||||
| TrEMBL | A0A078IU52 | 0.0 | A0A078IU52_BRANA; BnaCnng27630D protein | ||||
| TrEMBL | A0A0D3CH36 | 0.0 | A0A0D3CH36_BRAOL; Uncharacterized protein | ||||
| TrEMBL | A0A3P6EKS8 | 0.0 | A0A3P6EKS8_BRAOL; Uncharacterized protein | ||||
| STRING | Bo5g094500.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1467 | 28 | 91 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22780.1 | 0.0 | Tesmin/TSO1-like CXC domain-containing protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



