 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G22780.1 | ||||||||
| Common Name | ATTSO1, MWI23.15, TSO1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 695aa MW: 76276.8 Da PI: 6.675 | ||||||||
| Description | Tesmin/TSO1-like CXC domain-containing protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 50.5 | 4e-16 | 399 | 437 | 2 | 40 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
++k+CnCkkskClk+YCeCfaag++C e C+C dC+Nk
AT3G22780.1 399 SCKRCNCKKSKCLKLYCECFAAGVYCIEPCSCIDCFNKP 437
689**********************************96 PP
| |||||||
| 2 | TCR | 49.7 | 7.4e-16 | 484 | 522 | 1 | 39 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNk 39
++k+gCnCkks+C+kkYCeC++ g+ Cs +C+Ce+C N
AT3G22780.1 484 RHKRGCNCKKSNCMKKYCECYQGGVGCSMNCRCEGCTNV 522
589**********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 2.3E-19 | 398 | 439 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 36.989 | 399 | 524 | IPR005172 | CRC domain |
| Pfam | PF03638 | 6.7E-12 | 401 | 436 | IPR005172 | CRC domain |
| SMART | SM01114 | 5.6E-16 | 484 | 525 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 7.0E-11 | 486 | 521 | IPR005172 | CRC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009934 | Biological Process | regulation of meristem structural organization | ||||
| GO:0048444 | Biological Process | floral organ morphogenesis | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 695 aa Download sequence Send to blast |
MDKSQKNPTS QIGTSTPKSK FEDSPVFNYI SNLSPIESVK SISTAQTFSS LSFTSPPPVF 60 TSPHVISHRE SRFFRCHNSV DRSKHLESLD GSAVKGEVVV PLVEDLNKEA SLEDEEETSV 120 ETSSELPQIM KFDSQTSEHS DSPCTEDVVI EASSDPPRGD NGSSSEDVTM GLQNMLVVRE 180 GNDTPGCGRL ISDATELLVF RSPNDSEAFR CLVDKISSSE RRFCAGVKST KRPDINKDIP 240 ANGSSNENQP LAVLPTNESV FNLHRGGMRR RCLDFEMPGK RKKDIVDDQQ SVCDNNVAGE 300 SSSSCVVPGI GLHLNAVAMS AKDSNISVIH GYSISGEIQK SFSGSTTPIQ SQDTVQETSD 360 QAENEPVEEV PKALVFPELN LGSLKKKMRK SEQAGEGESC KRCNCKKSKC LKLYCECFAA 420 GVYCIEPCSC IDCFNKPIHE ETVLATRKQI ESRNPLAFAP KVIRNADSIM EASDDASKTP 480 ASARHKRGCN CKKSNCMKKY CECYQGGVGC SMNCRCEGCT NVFGRKDGSL LVIMESKLEE 540 NQETYEKRIA KIQHNVEVSK EVEQNPSSDQ PSTPLPPYRH LVVHQPFLSK NRLPPTQFFL 600 GTGSSSFRKP NSDLAQSQNE KKPLETVTED KTEIMPEILL NSPIANIKAI SPNSKRVSPP 660 QPGSSESGSI LRRRGNGRKL ILRSIPAFPS LNPNQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 1e-15 | 403 | 522 | 14 | 121 | Protein lin-54 homolog |
| 5fd3_B | 1e-15 | 403 | 522 | 14 | 121 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 268 | 282 | RRRCLDFEMPGKRKK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.24953 | 0.0 | flower| leaf| root| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 6739820 | 0.0 | ||||
| Genevisible | 258324_at | 0.0 | ||||
| Expression Atlas | AT3G22780 | - | ||||
| AtGenExpress | AT3G22780 | - | ||||
| ATTED-II | AT3G22780 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed at the first stage of pollen development in uninucleate microspores and bicellular pollen but not in tricellular and mature pollen. {ECO:0000269|PubMed:18057042}. | |||||
| Uniprot | TISSUE SPECIFICITY: Ubiquitous but expressed mostly in flowers with highest levels in developing ovules and microspores. {ECO:0000269|PubMed:10769244, ECO:0000269|PubMed:10769245, ECO:0000269|PubMed:18057042}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | putative DNA binding protein (tso1) mRNA, tso1-3 allele, | |||||
| UniProt | Probable floral-specific cell division component, required for proper organ formation in flowers. Regulates the floral meristem cell division and the inflorescence meristem organization. Plays a role in development of both male and female reproductive tissues. {ECO:0000269|PubMed:10769244, ECO:0000269|PubMed:10769245, ECO:0000269|PubMed:9043081, ECO:0000269|PubMed:9725857}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
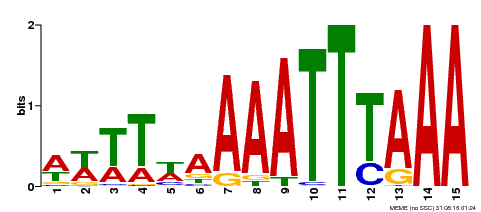
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00370 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G22780.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Despite a wild-type flower phenotype, the null mutant presents defects in pollen, carpel and ovule development, including a failure in megasporogenesis. Small siliques. {ECO:0000269|PubMed:18057042, ECO:0000269|PubMed:22072982, ECO:0000269|PubMed:9725857}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G22780 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF204059 | 0.0 | AF204059.1 Arabidopsis thaliana CXC domain protein TSO1 (TSO1) mRNA, complete cds. | |||
| GenBank | AF206324 | 0.0 | AF206324.1 Arabidopsis thaliana putative DNA binding protein (tso1) mRNA, tso1-3 allele, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_566718.2 | 0.0 | Tesmin/TSO1-like CXC domain-containing protein | ||||
| Swissprot | Q9LUI3 | 0.0 | TSO1_ARATH; CRC domain-containing protein TSO1 | ||||
| TrEMBL | A0A178VBV1 | 0.0 | A0A178VBV1_ARATH; TSO1 | ||||
| STRING | AT3G22780.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1467 | 28 | 91 | Representative plant | OGRP993 | 17 | 55 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G22780.1 |
| Entrez Gene | 821849 |
| iHOP | AT3G22780 |
| wikigenes | AT3G22780 |



