 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Rsa1.0_01007.1_g00004.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 475aa MW: 53587.4 Da PI: 7.8874 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.1 | 7.3e-17 | 230 | 276 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g WT +Ed ll ++v ++G+++W+ Ia++++ gR +kqc++rw+++l
Rsa1.0_01007.1_g00004.1 230 KGQWTSDEDRLLMQLVGLHGTKRWSQIAKMLQ-GRVGKQCRERWHNHL 276
799*****************************.*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 52.5 | 1.1e-16 | 283 | 324 | 2 | 45 |
SSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
+ W+++Ed +l++a+k+ G++ W+ Iar+++ gRt++ +k++w+
Rsa1.0_01007.1_g00004.1 283 DGWSEQEDRILIEAHKEIGNR-WAEIARKLP-GRTENTIKNHWN 324
56*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 29.077 | 225 | 280 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.06E-30 | 227 | 323 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.7E-15 | 229 | 278 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-27 | 231 | 284 | IPR009057 | Homeodomain-like |
| Pfam | PF13921 | 1.7E-17 | 233 | 290 | No hit | No description |
| CDD | cd00167 | 4.77E-14 | 233 | 276 | No hit | No description |
| PROSITE profile | PS51294 | 19.999 | 281 | 331 | IPR017930 | Myb domain |
| SMART | SM00717 | 8.9E-16 | 281 | 329 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.03E-12 | 285 | 327 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.8E-20 | 285 | 329 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010439 | Biological Process | regulation of glucosinolate biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1904095 | Biological Process | negative regulation of endosperm development | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 475 aa Download sequence Send to blast |
MHYPYLASVI FDDNVTLKDV HPSLTDDASF VQNVHHKPFH ANLDRSHVPS IINSDLCHQN 60 PILKISNATS DHIPNLNISP RHYDMQPHTF DSVSKYIFNG ITSSPCTEAF EAYLRGISSD 120 QDLFNMACNT SHPVESNDLP NISHVPIDNT MWDNNRNQGI IFGTESIFNL AMVDSYGAPK 180 PICILGANDD TTMNQHQNNQ IMIKADQIKK NKRFQMRRGS KLTRKTSIIK GQWTSDEDRL 240 LMQLVGLHGT KRWSQIAKML QGRVGKQCRE RWHNHLRPDI KKDGWSEQED RILIEAHKEI 300 GNRWAEIARK LPGRTENTIK NHWNATKRKQ HSRKFKGKDE VSLALGSDTL QNYIRSVTYN 360 DDTFMTTTSK SNTNNGRKNM RGKGKAVIVS ASGYDKIDDD NECTYIVDGV MNLNLDKSAT 420 CVSKSTSASG STVGSGSGVT EVDEPMADSW MVMHGCDEVM MKEIALLEMI AHGRI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 7e-45 | 228 | 330 | 2 | 104 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes the motif 5'-TAACGG-3' in the promoter of endosperm-induced genes (PubMed:27681170, PubMed:25194028, PubMed:19066902). Promotes vegetative-to-embryonic transition and the formation of somatic embryos from root explants in a WUS-independent manner but via the expression of embryonic genes (e.g. LEC1, LEC2, FUS3 and WUS) (PubMed:18695688). May play an important role during embryogenesis and seed maturation (PubMed:19066902, PubMed:25194028). Together with MYB115, activates the transcription of S-ACP-DES2/AAD2 and S-ACP-DES3/AAD3 thus promoting the biosynthesis of omega-7 monounsaturated fatty acid in seed endosperm (PubMed:27681170). Regulates negatively maturation genes in the endosperm (PubMed:25194028). {ECO:0000269|PubMed:18695688, ECO:0000269|PubMed:19066902, ECO:0000269|PubMed:25194028, ECO:0000269|PubMed:27681170}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00385 | DAP | Transfer from AT3G27785 | Download |
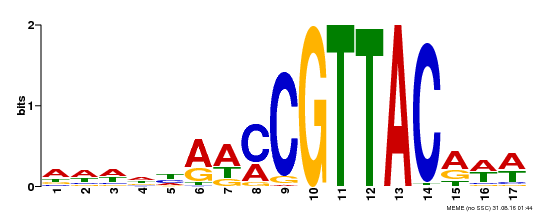
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Rsa1.0_01007.1_g00004.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by LEC2. {ECO:0000269|PubMed:25194028}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018438673.1 | 0.0 | PREDICTED: myb-like protein Q | ||||
| Swissprot | Q9LVW4 | 0.0 | MY118_ARATH; Transcription factor MYB118 | ||||
| TrEMBL | A0A3P6AHG5 | 0.0 | A0A3P6AHG5_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4EW44 | 0.0 | M4EW44_BRARP; Uncharacterized protein | ||||
| STRING | Bra033027.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6052 | 26 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27785.1 | 0.0 | myb domain protein 118 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



