 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G27785.1 | ||||||||
| Common Name | ATMYB118, MYB118, PGA37 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 437aa MW: 49492.1 Da PI: 7.6888 | ||||||||
| Description | myb domain protein 118 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 60.9 | 2.7e-19 | 189 | 235 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g WT+eEd+llv++v ++G++ W+ Ia++++ gR +kqc++rw+++l
AT3G27785.1 189 KGQWTPEEDKLLVQLVDLHGTKKWSQIAKMLQ-GRVGKQCRERWHNHL 235
799*****************************.*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 55.5 | 1.3e-17 | 242 | 283 | 2 | 45 |
SSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
+ WT+eEd +l++a+k+ G++ W+ Iar+++ gRt++ +k++w+
AT3G27785.1 242 DGWTEEEDIILIKAHKEIGNR-WAEIARKLP-GRTENTIKNHWN 283
56*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 30.877 | 184 | 239 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.95E-32 | 186 | 282 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.2E-17 | 188 | 237 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.3E-28 | 190 | 241 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.18E-15 | 192 | 235 | No hit | No description |
| Pfam | PF13921 | 6.5E-20 | 192 | 250 | No hit | No description |
| SMART | SM00717 | 3.1E-16 | 240 | 288 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.737 | 240 | 290 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-21 | 242 | 288 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.24E-12 | 244 | 286 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006357 | Biological Process | regulation of transcription from RNA polymerase II promoter | ||||
| GO:0010439 | Biological Process | regulation of glucosinolate biosynthetic process | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1904095 | Biological Process | negative regulation of endosperm development | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000981 | Molecular Function | RNA polymerase II transcription factor activity, sequence-specific DNA binding | ||||
| GO:0001135 | Molecular Function | transcription factor activity, RNA polymerase II transcription factor recruiting | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000014 | anatomy | rosette leaf | ||||
| PO:0009001 | anatomy | fruit | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009046 | anatomy | flower | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 437 aa Download sequence Send to blast |
MEFESVFKMH YPYLAAVIYD DSSTLKDFHP SLTDDFSCVH NVHHKPSMPH TYEIPSKETI 60 RGITPSPCTE AFEACFHGTS NDHVFFGMAY TTPPTIEPNV SHVSHDNTMW ENDQNQGFIF 120 GTESTLNQAM ADSNQFNMPK PLLSANEDTI MNRRQNNQVM IKTEQIKKKN KRFQMRRICK 180 PTKKASIIKG QWTPEEDKLL VQLVDLHGTK KWSQIAKMLQ GRVGKQCRER WHNHLRPDIK 240 KDGWTEEEDI ILIKAHKEIG NRWAEIARKL PGRTENTIKN HWNATKRRQH SRRTKGKDEI 300 SLSLGSNTLQ NYIRSVTYND DPFMTANANA NIGPRNMRGK GKNVMVAVSE YDEGECKYIV 360 DGVNNLGLED GRIKMPSLAA MSASGSASTS GSASGSGSGV TMEIDEPMTD SWMVMHGCDE 420 VMMNEIALLE MIAHGRL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 3e-44 | 187 | 289 | 2 | 104 | MYB PROTO-ONCOGENE PROTEIN |
| 1mse_C | 3e-44 | 187 | 289 | 2 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 3e-44 | 187 | 289 | 2 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.8032 | 0.0 | seed | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 15375296 | 0.0 | ||||
| Genevisible | 258233_at | 0.0 | ||||
| Expression Atlas | AT3G27785 | - | ||||
| AtGenExpress | AT3G27785 | - | ||||
| ATTED-II | AT3G27785 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed in embryos from the early heart stage and throughout embryogenesis (PubMed:18695688, PubMed:19066902). Induced at the onset of the maturation phase in the endosperm, in a high and homogeneous repartition (PubMed:18695688, PubMed:19066902, PubMed:25194028, PubMed:27681170). {ECO:0000269|PubMed:18695688, ECO:0000269|PubMed:19066902, ECO:0000269|PubMed:25194028, ECO:0000269|PubMed:27681170}. | |||||
| Uniprot | TISSUE SPECIFICITY: Mainly expressed in siliques (PubMed:18695688, PubMed:19066902). Also detected at low levels in leaves and flowers (PubMed:19066902). {ECO:0000269|PubMed:18695688, ECO:0000269|PubMed:19066902}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | putative transcription factor (MYB118) | |||||
| UniProt | Transcription activator that recognizes the motif 5'-TAACGG-3' in the promoter of endosperm-induced genes (PubMed:27681170, PubMed:25194028, PubMed:19066902). Promotes vegetative-to-embryonic transition and the formation of somatic embryos from root explants in a WUS-independent manner but via the expression of embryonic genes (e.g. LEC1, LEC2, FUS3 and WUS) (PubMed:18695688). May play an important role during embryogenesis and seed maturation (PubMed:19066902, PubMed:25194028). Together with MYB115, activates the transcription of S-ACP-DES2/AAD2 and S-ACP-DES3/AAD3 thus promoting the biosynthesis of omega-7 monounsaturated fatty acid in seed endosperm (PubMed:27681170). Regulates negatively maturation genes in the endosperm (PubMed:25194028). {ECO:0000269|PubMed:18695688, ECO:0000269|PubMed:19066902, ECO:0000269|PubMed:25194028, ECO:0000269|PubMed:27681170}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00385 | DAP | 27203113 | Download |
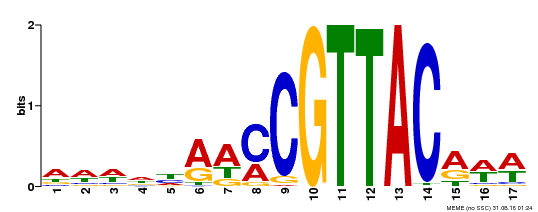
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G27785.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by LEC2. {ECO:0000269|PubMed:25194028}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- ATRM (Manually Curated Target Genes) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Target Gene (A: Activate/R: Repress) | |||||
| ATRM | AT1G21970(A) | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G27785 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ446708 | 0.0 | DQ446708.1 Arabidopsis thaliana clone pENTR221-At3g27785 myb family transcription factor (At3g27785) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_189416.2 | 0.0 | myb domain protein 118 | ||||
| Swissprot | Q9LVW4 | 0.0 | MY118_ARATH; Transcription factor MYB118 | ||||
| TrEMBL | A0A178VHH6 | 0.0 | A0A178VHH6_ARATH; PGA37 | ||||
| STRING | AT3G27785.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6052 | 26 | 43 | Representative plant | OGRP5 | 17 | 1784 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G27785.1 |
| Entrez Gene | 822399 |
| iHOP | AT3G27785 |
| wikigenes | AT3G27785 |



