 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | RrC10366_p1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Raphanus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1321aa MW: 147399 Da PI: 7.2448 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.2 | 0.00054 | 1204 | 1229 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
ykC C++sFs+ +L H r+ +
RrC10366_p1 1204 YKCDieGCTMSFSSEKQLSMHKRNiC 1229
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 14.9 | 7.5e-05 | 1229 | 1251 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp CgksF ++ +L++H r+H
RrC10366_p1 1229 CPvkGCGKSFFSHKYLVQHRRVH 1251
9999*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.1E-16 | 17 | 58 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.689 | 18 | 59 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 7.5E-14 | 19 | 52 | IPR003349 | JmjN domain |
| SuperFamily | SSF51197 | 1.84E-26 | 129 | 176 | No hit | No description |
| SMART | SM00558 | 1.4E-53 | 198 | 367 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 34.235 | 201 | 367 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.84E-26 | 216 | 382 | No hit | No description |
| Pfam | PF02373 | 3.8E-38 | 231 | 350 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 9.1E-4 | 1203 | 1226 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 7.1 | 1204 | 1226 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.214 | 1227 | 1256 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0027 | 1227 | 1251 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-6 | 1228 | 1255 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1229 | 1251 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.5E-10 | 1243 | 1285 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 8.2E-10 | 1256 | 1280 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.801 | 1257 | 1286 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0015 | 1257 | 1281 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1259 | 1281 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.42E-8 | 1275 | 1309 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.9E-9 | 1281 | 1309 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.157 | 1287 | 1318 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.2 | 1287 | 1313 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1289 | 1313 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1321 aa Download sequence Send to blast |
MSEQSTDVFP WLKSLPVAPE FRPTLAEFQD PIAYIFKIEE EASRYGICKI LPPLPPPSKK 60 TTVSNLNRSL AARASTHDGA CDYDGSPAFA TRQQQVGFCP RKQRPVQRPV WQSGERYSFG 120 EFEAKTKTFE KSYLKRCCGG GRKSQQLSAL EIETLYWRAT VDKPFTVEYA NDMPGSAFIP 180 LSLAAARRRE YGGSDGGTVG ESAWNMRAMA RAEGSLLRFM KEEIPGVTSP MVYVAMMFSW 240 FAWHVEDHDL HSLNYLHMGA GKTWYGVPKD AALAFEEVVR DHGYGGELNP LVTFSTLGEK 300 TTVMSPEVFV KAGIPCCRLV QNPGEFVVTF PRAYHSGFSH GFNCGEASNI ATPEWLRVAK 360 EAAIRRAAIS YPPMVSHLQL LYDYALALGS RVPASIHAKP RSSRLKDKKR SEGERLSKEL 420 FAQNVIHNNE LLCSLGKGSP VALLPQSSSD VSVCSDLRIG SHLSANQEKP IQPKSEDLSP 480 ESNTVCLKET VSVKEKFTSL CGRNINQETM PEAERREKDG AVGLSDQRLF SCVSCGIICF 540 DCVAIVQPKE AAARYLSSAD CSFFSDRTVS SGSANPGQDE RSLVVEPHPQ SMGKRDVGCI 600 YSAPVQSTDH SIMTIDQGTS TASSALGLLA SAYGDSSDSE EEEHKGIDAV ISQNGASSYK 660 TPAFCTDGNE EATADGGTSD HSSERLSCGK GKGVDEGSNG GSASVLKVTL PFVSSSDDDS 720 SRLHVFCLEH AAEVEQQLRP FGGINIMLLC HPEYPRIEAE GKVVAEELGV KHEWSDTEFR 780 NVTREDEETI QAALVNVEAK AGNRDWTVKL SINLSYSAIL SRSPLYSKQM PYNSVIYNMF 840 GRSSPSTSSP TQHQVSGQRS CRQRKYVVGK WCGKVWMSHQ VHPFLLQEDS EGEETERSHL 900 RVEDVSGERL SPCDVSRDAT TMFGRKYSRK RKMRAKAVTR KKLTSFKRED GVSDDTSEDH 960 SYKQQWRRGF GNEEEMSDDS SDPMSDHQQQ QLLKGIRRDK GVREFESDDE VSDRSLGEEY 1020 AARECADSDS STDIDALEDS VYDGHNDDSI HKHPRSKRAK VIRIPVYQQR GNIQASQIGG 1080 DYDSEDDSLE EQEGFSTRKR QTRSTAKRKV KTEIVESPRD TRGRRTLQQS GYRKKIQDLD 1140 PYMEGPSTRL RVRNLKPSRK SSETKPKRGS SNASFSRVVS EEEELEEDDE EEGEEERAAA 1200 ASAYKCDIEG CTMSFSSEKQ LSMHKRNICP VKGCGKSFFS HKYLVQHRRV HSDDRPLKCP 1260 WKGCKMTFKW AWSRTEHIRV HTGERPYHCA EPGCGQTFRF VSDFSRHKKK TGHSAKKTKK 1320 R |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 3e-75 | 6 | 390 | 3 | 355 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 3e-75 | 6 | 390 | 3 | 355 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 927 | 932 | SRKRKM |
| 2 | 929 | 941 | KRKMRAKAVTRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
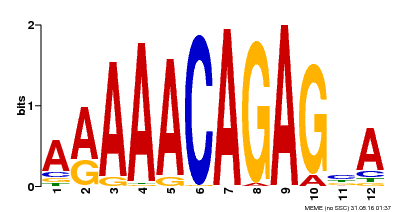
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018458317.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6-like isoform X1 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | V4M0K3 | 0.0 | V4M0K3_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006404260.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5259 | 28 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



