 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.010G114800.1 | ||||||||
| Common Name | PHAVU_010G114800g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 631aa MW: 69071.7 Da PI: 6.9111 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40.4 | 5.3e-13 | 464 | 510 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI++Lq
Phvul.010G114800.1 464 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAYINELQ 510
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.6E-52 | 51 | 241 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.047 | 460 | 509 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.48E-15 | 463 | 514 | No hit | No description |
| SuperFamily | SSF47459 | 2.49E-18 | 463 | 523 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.9E-10 | 464 | 510 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 3.2E-18 | 464 | 523 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.2E-17 | 466 | 515 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 631 aa Download sequence Send to blast |
MKIEVGLGGV WNEEEKATVV EVLGRGAFDY LVANSVSNEN LLMAFGSGEN LQNKLSDLVE 60 RPNASNFSWN YAIFWQISQS KFGDWVLGWG DGCCREPREG EEGGEVRGKG RGVVADDEKV 120 QGMRKRVLQK LHMTFGGSDE DNYAFGLDRV TDTEMFFLAS MYFSFPRGLG GPGKCFASGK 180 HLWLSDVFKS SFDYCVRSFL AKSAGIQTVV LVPTDMGVVE MGSVRMVGES FELLQALKSV 240 FSSQASLVPP RVKPIAPLDL VSEKREENAD APFSSVPIGD KDNNNSSNGN NRVEGIGVPK 300 IFGQDLNSVT QFREKLAVRK MEERPRAWGG HPNGNTIGFP NGIHTSNWGT GQVVRQVAPP 360 EMHARRLASS GTLSVPVPEL ANGTRHDFVH NNYQQQRPVQ MQIDFSGATS RGATSRASGR 420 SIIAESEISD VEASCKEERV SVADDRRPRK RGRKPANGRE EPLNHVEAER QRREKLNQRF 480 YALRAVVPNI SKMDKASLLG DAIAYINELQ AKLKTMESER ERFGSTSMDG SAVEANARAE 540 NHSNGAPDVD VQAAQDGVVV KVSCPIDVHP VSKVIQTFKE AEIGVVDSKL TAANDTVFHT 600 FVVKSQGPDQ LTKEKLIALF SKASNPIQTS * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 4e-30 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 4e-30 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 4e-30 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 4e-30 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 4e-30 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 4e-30 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 4e-30 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 4e-30 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 445 | 453 | RRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
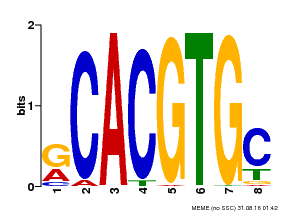
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.010G114800.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015042 | 0.0 | AP015042.1 Vigna angularis var. angularis DNA, chromosome 9, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007135266.1 | 0.0 | hypothetical protein PHAVU_010G114800g | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | V7APM6 | 0.0 | V7APM6_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007135266.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5448 | 32 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 1e-165 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.010G114800.1 |
| Entrez Gene | 18618140 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



