 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.007G195900.1 | ||||||||
| Common Name | PHAVU_007G195900g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | NF-YB | ||||||||
| Protein Properties | Length: 146aa MW: 16272.4 Da PI: 8.0596 | ||||||||
| Description | NF-YB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NF-YB | 158.8 | 8.4e-50 | 4 | 99 | 2 | 97 |
NF-YB 2 reqdrflPianvsrimkkvlPanakiskdaketvqecvsefisfvtseasdkcqrekrktingddllwalatlGfedyveplkvylkkyrele 94
+eqd+ lPianv+rimk++lP akisk+ k+++qecv+efisfvt+easdkc++e+rkt+ngdd++wal++lGf++y+e++ yl+kyr++e
Phvul.007G195900.1 4 EEQDKALPIANVGRIMKQILPPSAKISKEGKQLMQECVTEFISFVTGEASDKCHKENRKTVNGDDICWALSSLGFDNYAETIARYLHKYRQAE 96
69******************************************************************************************* PP
NF-YB 95 gek 97
+ek
Phvul.007G195900.1 97 REK 99
997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.20.10 | 2.3E-48 | 3 | 111 | IPR009072 | Histone-fold |
| SuperFamily | SSF47113 | 2.48E-37 | 6 | 111 | IPR009072 | Histone-fold |
| Pfam | PF00808 | 2.6E-25 | 10 | 73 | IPR003958 | Transcription factor CBF/NF-Y/archaeal histone domain |
| PRINTS | PR00615 | 9.5E-19 | 37 | 55 | No hit | No description |
| PROSITE pattern | PS00685 | 0 | 40 | 56 | IPR003956 | Transcription factor, NFYB/HAP3, conserved site |
| PRINTS | PR00615 | 9.5E-19 | 56 | 74 | No hit | No description |
| PRINTS | PR00615 | 9.5E-19 | 75 | 93 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 146 aa Download sequence Send to blast |
MDQEEQDKAL PIANVGRIMK QILPPSAKIS KEGKQLMQEC VTEFISFVTG EASDKCHKEN 60 RKTVNGDDIC WALSSLGFDN YAETIARYLH KYRQAEREKM NHNKKYEGSA TQTLNQPQLI 120 LPPPPPSLSR VTGKVQNPPT TDQSG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4g91_B | 5e-41 | 5 | 94 | 3 | 92 | Transcription factor HapC (Eurofung) |
| 4g92_B | 5e-41 | 5 | 94 | 3 | 92 | Transcription factor HapC (Eurofung) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Component of the NF-Y/HAP transcription factor complex. The NF-Y complex stimulates the transcription of various genes by recognizing and binding to a CCAAT motif in promoters. | |||||
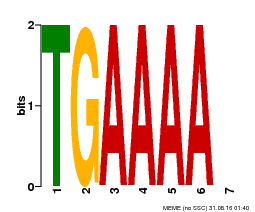
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00133 | DAP | Transfer from AT1G09030 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.007G195900.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015041 | 1e-176 | AP015041.1 Vigna angularis var. angularis DNA, chromosome 8, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007144936.1 | 1e-107 | hypothetical protein PHAVU_007G195900g | ||||
| Swissprot | O04027 | 3e-54 | NFYB4_ARATH; Nuclear transcription factor Y subunit B-4 | ||||
| TrEMBL | V7BHD2 | 1e-105 | V7BHD2_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007144936.1 | 1e-106 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF805 | 32 | 131 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G09030.1 | 3e-56 | nuclear factor Y, subunit B4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.007G195900.1 |
| Entrez Gene | 18626423 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



