 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G09030.1 | ||||||||
| Common Name | F7G19.10, HAP3D, NFYB4, NF-YB4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | NF-YB | ||||||||
| Protein Properties | Length: 139aa MW: 15741.4 Da PI: 7.5093 | ||||||||
| Description | nuclear factor Y, subunit B4 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NF-YB | 163.4 | 3e-51 | 2 | 97 | 2 | 97 |
NF-YB 2 reqdrflPianvsrimkkvlPanakiskdaketvqecvsefisfvtseasdkcqrekrktingddllwalatlGfedyveplkvylkkyrelegek 97
+++dr+lPianv+r+mk++lP+nakisk+ak+tvqec++efisfvt+eas+kc+re+rkt+ngdd++wal+tlG+++y++++ +l+kyre+e+e+
AT1G09030.1 2 TDEDRLLPIANVGRLMKQILPSNAKISKEAKQTVQECATEFISFVTCEASEKCHRENRKTVNGDDIWWALSTLGLDNYADAVGRHLHKYREAERER 97
689*******************************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.20.10 | 5.8E-49 | 2 | 123 | IPR009072 | Histone-fold |
| SuperFamily | SSF47113 | 4.33E-39 | 4 | 118 | IPR009072 | Histone-fold |
| Pfam | PF00808 | 5.2E-26 | 7 | 71 | IPR003958 | Transcription factor CBF/NF-Y/archaeal histone domain |
| PRINTS | PR00615 | 3.2E-16 | 35 | 53 | No hit | No description |
| PROSITE pattern | PS00685 | 0 | 38 | 54 | IPR003956 | Transcription factor, NFYB/HAP3, conserved site |
| PRINTS | PR00615 | 3.2E-16 | 54 | 72 | No hit | No description |
| PRINTS | PR00615 | 3.2E-16 | 73 | 91 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 139 aa Download sequence Send to blast |
MTDEDRLLPI ANVGRLMKQI LPSNAKISKE AKQTVQECAT EFISFVTCEA SEKCHRENRK 60 TVNGDDIWWA LSTLGLDNYA DAVGRHLHKY REAERERTEH NKGSNDSGNE KETNTRSDVQ 120 NQSTKFIRVV EKGSSSSAR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4g91_B | 3e-40 | 1 | 92 | 1 | 92 | Transcription factor HapC (Eurofung) |
| 4g92_B | 3e-40 | 1 | 92 | 1 | 92 | Transcription factor HapC (Eurofung) |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 264642_at | 0.0 | ||||
| Expression Atlas | AT1G09030 | - | ||||
| AtGenExpress | AT1G09030 | - | ||||
| ATTED-II | AT1G09030 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in flowers, siliques and young rosettes. {ECO:0000269|PubMed:11250072}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Component of the NF-Y/HAP transcription factor complex. The NF-Y complex stimulates the transcription of various genes by recognizing and binding to a CCAAT motif in promoters. | |||||
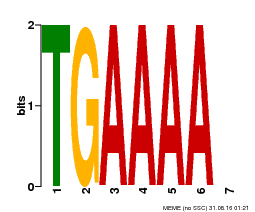
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00133 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G09030.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| BioGRID | AT1G54830, AT1G56170 | |||||
| IntAct | Search O04027 | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G09030 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT029363 | 0.0 | BT029363.1 Arabidopsis thaliana At1g09030 mRNA, complete cds. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| GenBank | F7G19 | 0.0 | AC000106.1 Sequence of BAC F7G19 from Arabidopsis thaliana chromosome 1, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_172377.1 | 1e-101 | nuclear factor Y, subunit B4 | ||||
| Swissprot | O04027 | 1e-102 | NFYB4_ARATH; Nuclear transcription factor Y subunit B-4 | ||||
| TrEMBL | A0A384L7L3 | 2e-99 | A0A384L7L3_ARATH; NF-YB4 | ||||
| TrEMBL | C0SUU0 | 2e-99 | C0SUU0_ARATH; Uncharacterized protein At1g09030 (Fragment) | ||||
| STRING | AT1G09030.1 | 1e-100 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM255 | 28 | 229 | Representative plant | OGRP168 | 17 | 170 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G09030.1 |
| Entrez Gene | 837424 |
| iHOP | AT1G09030 |
| wikigenes | AT1G09030 |



