 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.9NG271400.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1005aa MW: 111651 Da PI: 9.3504 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 10.9 | 0.0014 | 911 | 933 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cg+ F ++ +L +H ++H
Pavir.9NG271400.2.p 911 CPvkGCGRKFFSHKYLLQHRKVH 933
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 11.7 | 0.00082 | 969 | 995 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+Cp C ++F+ s++ rH r+ H
Pavir.9NG271400.2.p 969 YVCPepGCEQTFRFVSDFSRHKRRtgH 995
99********************98666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 3.3E-4 | 879 | 908 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 14 | 886 | 908 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 1.1 | 909 | 933 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.905 | 909 | 938 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.89E-5 | 910 | 945 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 911 | 933 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 6.6E-4 | 911 | 937 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-9 | 938 | 962 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 9.7E-4 | 939 | 963 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.718 | 939 | 968 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 941 | 963 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.82E-10 | 949 | 993 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-10 | 963 | 991 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.17 | 969 | 995 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.367 | 969 | 1000 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 971 | 995 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1005 aa Download sequence Send to blast |
MATPEWLRVA KEAAVRRASI NRPPMVSHYQ LLYELALSMC LRDPSSGAVE PRSSRLKEKK 60 KGEGEQLVKK IFVQNVIEDN KLLNHFLRDG SSCIILPTSP NDGSALSTLL SKSQSTTKSR 120 ISDCHCNITE APKDSRCLPV NGLLGKNGEL SSSKELSTSV CSGEKFLPTA CMHDCVNMSG 180 SSDANNAESD KGDINSTAGL LDQGFLSCVT CGILSFSCVA VIKPRECAAK WLMSADSSLI 240 NKQFADSGES HLIDALQSAT TNSGILRSGF EMDGNRTSSG AAALNRNSAL DLLASAYGDP 300 SDSDEDVLNK KNQVANVSSE LISRTIQSQP NNISTIGGCD GTNLSSSSKE HQQGPSSQRS 360 QCIGNNNNGP KGVRTRNKYQ LKMVLSEGFQ PKDKYSEIQK KVQCEPLSSN KASTEQLRGT 420 DCQAGHNSAT ICMDSNTSTT TRVDNLATSI VKPDKDSSRM HVFCLEHAIE VEKQLQTIGG 480 AHIFLLCRPE YPKIEVEAKL LAEEVEVEYD WKDIRFKETS IEDRKKMREV VQDEETIPTN 540 SDWAVKMGIN LYYSANLAKS PLYNKQLPYN RVIYKAFGCS SPNNSPVKLK TYSRRQGRAK 600 KIVLAGRWCG KVWMSNQVHP YLAHRIKSHE PDEINEICSS VQKSNSEHVE KSSREATCTR 660 KSNSRAIEEK TSKREEEPLE KANANTPKHS EEDNSKALEG AAEASAIKSS SRTVVEETSK 720 RKKEPLEKAN TKKPKHTEEE NSKALKGASE ASHPSPSRMV IRSSSRIANR KNMLKSKMEE 780 KDNDPANRPK AMVEEDGNDP ASCSRARPLR QKTNTDVKKQ TKKPRAEKQK APSPTALEDE 840 DKELSFTKQQ LSSRKQKTKV EEKQQMKKTR ENKGAPPSSP KQGEEYACNI EGCSMSFGTK 900 QELSLHKRDI CPVKGCGRKF FSHKYLLQHR KVHNDDRPLK CSWKGCDMAF KWPWARTEHM 960 RVHTGDRPYV CPEPGCEQTF RFVSDFSRHK RRTGHAAKKA KTKK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 2e-60 | 886 | 999 | 23 | 136 | Lysine-specific demethylase REF6 |
| 6a58_A | 2e-60 | 886 | 999 | 23 | 136 | Lysine-specific demethylase REF6 |
| 6a59_A | 2e-60 | 886 | 999 | 23 | 136 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 719 | 734 | KRKKEPLEKANTKKPK |
| 2 | 989 | 1003 | KRRTGHAAKKAKTKK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.26361 | 0.0 | callus | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaves and flag leaves. Expressed at low levels in roots, shoots, stems and panicles. {ECO:0000269|PubMed:24280387}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3 with a specific activity for H3K27me3 and H3K27me2. No activity on H3K4me3, H3K9me3, H3K27me1 and H3K36me3. Involved in biotic stress response. May demethylate H3K27me3-marked defense-related genes and increase their basal and induced expression levels during pathogen infection. {ECO:0000269|PubMed:24280387}. | |||||
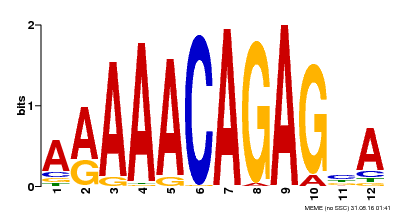
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.9NG271400.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt stress, abscisic acid (ABA), jasmonic acid (JA), the ethylene precursor ACC and infection by the bacterial pathogen Xanthomonas oryzae pv. oryzae. {ECO:0000269|PubMed:24280387}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025815151.1 | 0.0 | lysine-specific demethylase JMJ705-like | ||||
| Swissprot | Q5N712 | 0.0 | JM705_ORYSJ; Lysine-specific demethylase JMJ705 | ||||
| TrEMBL | A0A3L6SIW8 | 0.0 | A0A3L6SIW8_PANMI; Uncharacterized protein | ||||
| STRING | Pavir.J31927.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 3e-51 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.9NG271400.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



