 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.6NG358100.3.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 526aa MW: 54325.7 Da PI: 9.579 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.4 | 1.2e-05 | 60 | 82 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ C+k F r nL+ H r H
Pavir.6NG358100.3.p 60 FVCEVCNKGFQREQNLQLHRRGH 82
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.3 | 0.00026 | 136 | 158 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC +C+k++ +s+ k H +t+
Pavir.6NG358100.3.p 136 WKCDKCNKRYAVQSDWKAHSKTC 158
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 1.38E-7 | 59 | 82 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 7.1E-6 | 59 | 82 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.094 | 60 | 82 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0041 | 60 | 82 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF12171 | 3.4E-5 | 60 | 82 | IPR022755 | Zinc finger, double-stranded RNA binding |
| PROSITE pattern | PS00028 | 0 | 62 | 82 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 300 | 101 | 131 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.8E-5 | 124 | 157 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.38E-7 | 131 | 156 | No hit | No description |
| SMART | SM00355 | 110 | 136 | 156 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 526 aa Download sequence Send to blast |
MAAAHLFGLG DAQMQPPLPQ QAAPPPPAAA PAPKKKRNQP DPDAEVIALS PKSLLATNRF 60 VCEVCNKGFQ REQNLQLHRR GHNLPWKLKQ KNPKETRRRV YLCPEPTCVH HDPSRALGDL 120 TGIKKHYCRK HGEKKWKCDK CNKRYAVQSD WKAHSKTCGT REYRCDCGTL FSRRDSFITH 180 RAFCDAMAQE SARAPPIGAG MYGSGGMALG LSGLAASQLH SFQDQAHSSA ATAIGGNPAA 240 QFEHLMPSSA GSPAFRGAQP ASSSSSPFYL GGAEDGHQSQ PGHTSLLHGK PAFHGLMQLP 300 EQHGQPGSNG LLNLGFFSGA SSGQDARLVF PNQFNGATGG NGRGDGSEHG NSGANTGSAA 360 IFSGNLMGNQ MAGGGGGFSS SLYNSTETVA PPQMSATALL QKAAQMGATT SGGGGGSVNS 420 LLRGLGSGGA LNGRPSGAAG FMAGESSSSR STSQAENESQ FRDLMNSLAA SGSGGFPGMD 480 DGKLSTRDFL GVSGGVVRSM GGAAGLPLRH GAAGIGMGSL DPEMK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 1e-33 | 132 | 193 | 3 | 64 | Zinc finger protein JACKDAW |
| 5b3h_F | 1e-33 | 132 | 193 | 3 | 64 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.4524 | 0.0 | callus| leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in leaf tissues. {ECO:0000305|PubMed:22898356}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | Transfer from AT2G02070 | Download |
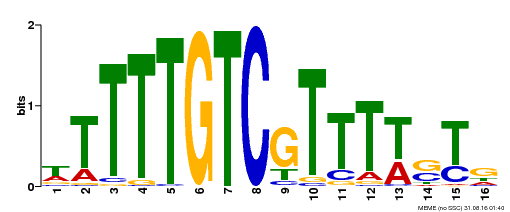
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.6NG358100.3.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT053822 | 0.0 | BT053822.1 Zea mays full-length cDNA clone ZM_BFb0209O02 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025821062.1 | 0.0 | protein indeterminate-domain 5, chloroplastic-like isoform X2 | ||||
| Swissprot | Q9ZUL3 | 1e-120 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | A0A3L6PM37 | 0.0 | A0A3L6PM37_PANMI; INDETERMINATE-related protein 1 | ||||
| STRING | Pavir.Fb02381.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G02080.1 | 1e-107 | indeterminate(ID)-domain 4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.6NG358100.3.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



