 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT2G02070.1 | ||||||||
| Common Name | AtIDD5, F5O4.16, IDD5, IDZ2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 602aa MW: 64463.7 Da PI: 9.3288 | ||||||||
| Description | indeterminate(ID)-domain 5 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.1 | 3.1e-05 | 81 | 103 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ C+k F r nL+ H r H
AT2G02070.1 81 FICEVCNKGFQREQNLQLHRRGH 103
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 12.5 | 0.00046 | 157 | 179 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC +C+k++ +s+ k H +t+
AT2G02070.1 157 WKCDKCSKRYAVQSDWKAHSKTC 179
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 6.8E-6 | 80 | 103 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.74E-7 | 80 | 103 | No hit | No description |
| Pfam | PF12171 | 3.0E-5 | 81 | 103 | IPR022755 | Zinc finger, double-stranded RNA binding |
| SMART | SM00355 | 0.0052 | 81 | 103 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.99 | 81 | 103 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 83 | 103 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 110 | 122 | 152 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 4.0E-5 | 145 | 178 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.74E-7 | 152 | 177 | No hit | No description |
| SMART | SM00355 | 140 | 157 | 177 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009507 | Cellular Component | chloroplast | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 602 aa Download sequence Send to blast |
MAASSSSAAS FFGVRQDDQS HLLPPNSSAA APPPPPPHHQ APLPPLEAPP QKKKRNQPRT 60 PNSDAEVIAL SPKTLMATNR FICEVCNKGF QREQNLQLHR RGHNLPWKLK QKSTKEVKRK 120 VYLCPEPSCV HHDPSRALGD LTGIKKHYYR KHGEKKWKCD KCSKRYAVQS DWKAHSKTCG 180 TKEYRCDCGT LFSRRDSFIT HRAFCDALAQ ESARHPTSLT SLPSHHFPYG QNTNNSNNNA 240 SSMILGLSHM GAPQNLDHQP GDVLRLGSGG GGGGAASRSS SDLIAANASG YFMQEQNPSF 300 HDQQDHHHHH QQGFLAGNNN IKQSPMSFQQ NLMQFSHDNH NSAPSNVFNL SFLSGNNGVT 360 SATSNPNAAA AAAVSSGNLM ISNHYDGENA VGGGGEGSTG LFPNNLMSSA DRISSGSVPS 420 LFSSSMQSPN SAPHMSATAL LQKAAQMGST SSNNNNGSNT NNNNNASSIL RSFGSGIYGE 480 NESNLQDLMN SFSNPGATGN VNGVDSPFGS YGGVNKGLSA DKQSMTRDFL GVGQIVKSMS 540 GSGGFQQQQQ QQQQQQQQQQ HGNSRERVGS SSDSADRSSM NVNTGGGPAS TSPPYGIHHA 600 SF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 2e-33 | 153 | 214 | 3 | 64 | Zinc finger protein JACKDAW |
| 5b3h_F | 2e-33 | 153 | 214 | 3 | 64 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.13356 | 0.0 | bud| flower| seed | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 266120_at | 0.0 | ||||
| Expression Atlas | AT2G02070 | - | ||||
| AtGenExpress | AT2G02070 | - | ||||
| ATTED-II | AT2G02070 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in leaf tissues. {ECO:0000305|PubMed:22898356}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | 27203113 | Download |
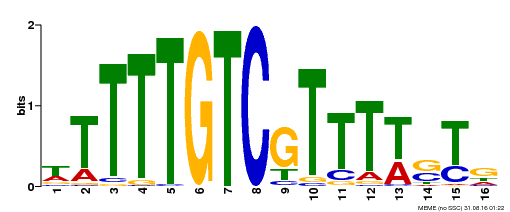
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT2G02070.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| BioGRID | AT1G50420 | |||||
| IntAct | Search Q9ZUL3 | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Down-regulation of SS4 during the light period of both short and long day conditions. Deformity of the chloroplasts and their contained starch granules, with an increased number of starch granules per chloroplast. {ECO:0000269|PubMed:22898356}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT2G02070 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ621495 | 0.0 | AJ621495.1 Arabidopsis thaliana mRNA for ID1-like zinc finger protein 2 (idz2 gene). | |||
| GenBank | AY056174 | 0.0 | AY056174.1 Arabidopsis thaliana putative C2H2-type zinc finger protein (At2g02070) mRNA, complete cds. | |||
| GenBank | BT001215 | 0.0 | BT001215.1 Arabidopsis thaliana putative C2H2-type zinc finger protein (At2g02070) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001324413.1 | 0.0 | indeterminate(ID)-domain 5 | ||||
| Refseq | NP_178316.1 | 0.0 | indeterminate(ID)-domain 5 | ||||
| Swissprot | Q9ZUL3 | 0.0 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | A0A178VZR3 | 0.0 | A0A178VZR3_ARATH; IDD5 | ||||
| STRING | AT2G02070.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4373 | 26 | 54 | Representative plant | OGRP95 | 16 | 242 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT2G02070.1 |
| Entrez Gene | 814738 |
| iHOP | AT2G02070 |
| wikigenes | AT2G02070 |



