 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.5NG079300.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 258aa MW: 27837.9 Da PI: 5.4466 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.2 | 9.9e-19 | 79 | 128 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
y+GVr++ +g+WvAeIr+p + r +r +lg+f ta eAa a+++a+k+++g
Pavir.5NG079300.1.p 79 AYRGVRQRT-WGKWVAEIREP---N-RgRRLWLGSFPTAVEAAHAYDEAAKAMYG 128
59****999.**********8...3.35*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 2.7E-13 | 78 | 128 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 21.641 | 79 | 136 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.44E-20 | 79 | 137 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 3.4E-34 | 79 | 142 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.8E-30 | 80 | 137 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.0E-9 | 80 | 91 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.26E-29 | 80 | 138 | No hit | No description |
| PRINTS | PR00367 | 4.0E-9 | 102 | 118 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 258 aa Download sequence Send to blast |
MDLGHGGQGG EGDSSGGQVR KKRARRKSTG PDSIAETIKW WKEQNQKLQD ESGSRKAPAK 60 GSKKGCMAGK GGPENGNCAY RGVRQRTWGK WVAEIREPNR GRRLWLGSFP TAVEAAHAYD 120 EAAKAMYGAK ARVNFSENSP DANSGCTSAL SLLASSVPAV ALHGFNEKDE VESVETEVHD 180 VKAQVNNDLG SIHVECTSVE VLQSGESVLQ KEGSVSYDYF NVEEVVEMII IELNADKKIE 240 VHEECLGGDD GFSLFAY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 1e-15 | 80 | 135 | 3 | 59 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.20564 | 0.0 | callus| leaf| root| stem | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity- and dehydration-inducible transcription (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00302 | DAP | Transfer from AT2G40340 | Download |
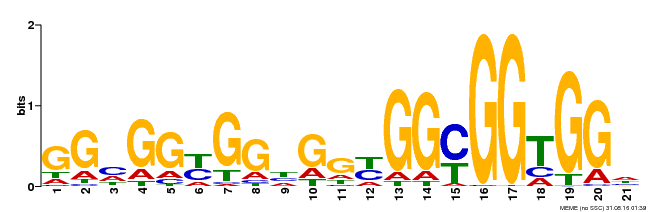
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.5NG079300.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HQ132744 | 0.0 | HQ132744.1 Setaria italica dehydration responsive element-binding protein 2 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025816059.1 | 0.0 | dehydration-responsive element-binding protein 2A-like | ||||
| Swissprot | A2WL19 | 1e-122 | DRE2A_ORYSI; Dehydration-responsive element-binding protein 2A | ||||
| TrEMBL | A0A3L6QX70 | 0.0 | A0A3L6QX70_PANMI; Dehydration-responsive element-binding protein 2A isoform X1 | ||||
| STRING | Pavir.J37967.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7315 | 35 | 51 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40340.1 | 1e-35 | ERF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.5NG079300.1.p |



