 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.5KG013700.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 473aa MW: 50915.9 Da PI: 7.5359 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 87.2 | 1.5e-27 | 197 | 251 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k +++WtpeLH+rFv+aveqL G++kA+P++il++m++++Lt+++++SHLQkYR++
Pavir.5KG013700.2.p 197 KRKVDWTPELHRRFVQAVEQL-GIDKAVPSRILDIMGIDCLTRHNIASHLQKYRSH 251
5799*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.638 | 194 | 253 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-26 | 195 | 254 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 7.35E-18 | 196 | 254 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-26 | 197 | 251 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.1E-6 | 200 | 249 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 473 aa Download sequence Send to blast |
MLEVSILRSP SKADHHQRGF VGDHVVFPTS GSVCCDDFMV DDNLLDYIDF SCDAVPFFDA 60 DGDILPDLEV DPTELLAEFS SPEASPTAAR AEAKPPDDLR RRQPEEEAVV LSTSTTTATI 120 VEEKKAAERV LSAKDEKLAA AGVSVTTRTM RKKVDGEGSS AEEDSGGAGS DTKWSASAGG 180 HGKKKTAAAD KNSSSGKRKV DWTPELHRRF VQAVEQLGID KAVPSRILDI MGIDCLTRHN 240 IASHLQKYRS HRKHLMAREA EAATWAQKRH QYMYAPAPAR KLDAAAAGGG PWVVPTTIGF 300 PRPALCRALH VWGHPPSAGG VDAPPTMLPV WPRHLAPPRP WAPPPAYWQH QYNAARKWGP 360 HHQAAAAAAA VTPAMVPQLP RFPVPPHPAT MYRPTCSSSM VVPPPPPTST KLADLQLQLD 420 AHPSKESIDA AIGDVLVKPW LPLPLGLKPP SMDSVMSELH KHGIPKVPPS SG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 3e-15 | 193 | 249 | 1 | 57 | ARR10-B |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.16046 | 1e-104 | leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaves. {ECO:0000269|PubMed:11340194}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that promotes chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
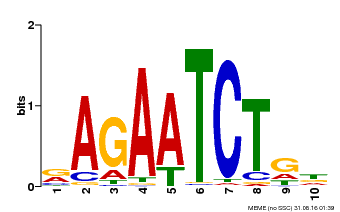
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.5KG013700.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. {ECO:0000269|PubMed:11340194}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF318579 | 1e-89 | AF318579.1 Zea mays putative transcription factor GOLDEN 2 mRNA, complete cds. | |||
| GenBank | BT062838 | 1e-89 | BT062838.1 Zea mays full-length cDNA clone ZM_BFb0374G22 mRNA, complete cds. | |||
| GenBank | EU956792 | 1e-89 | EU956792.1 Zea mays clone 1574106 transcription factor ZmGLK1 mRNA, complete cds. | |||
| GenBank | KJ727689 | 1e-89 | KJ727689.1 Zea mays clone pUT5628 G2-like transcription factor (GLK13) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025818980.1 | 0.0 | probable transcription factor GLK2 isoform X3 | ||||
| Swissprot | Q5NAN5 | 1e-102 | GLK2_ORYSJ; Probable transcription factor GLK2 | ||||
| TrEMBL | A0A2T8IPG3 | 0.0 | A0A2T8IPG3_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Ea00014.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 1e-54 | GBF's pro-rich region-interacting factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.5KG013700.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



