 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.3NG212900.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 270aa MW: 28762.2 Da PI: 8.8946 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 143.3 | 1.4e-44 | 14 | 150 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdL.pkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkd 89
lppGfrF+Ptdeelvv+yL++++ +++l+++ i++v+i+ ++P+dL p +e+e yfF++++ ky +g+r+nrat +gyWkatgk+
Pavir.3NG212900.1.p 14 LPPGFRFRPTDEELVVHYLRRRALASPLPAAVDIPDVRILAHDPSDLlP--PGWSEQERYFFTCKEAKYVKGRRANRATGAGYWKATGKE 101
79***************************9666**************43..3447899******************************** PP
NAM 90 kevlsk..........kgelvglkktLvfykgrapkgektdWvmheyrl 128
+v ++ + lvg+k++Lvfy+g+ p+g+ktdWvmheyrl
Pavir.3NG212900.1.p 102 MPVAVAvparggggqaQAVLVGMKRSLVFYRGKPPTGSKTDWVMHEYRL 150
***99989988865544455***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 3.01E-49 | 8 | 177 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 48.17 | 14 | 177 | IPR003441 | NAC domain |
| Pfam | PF02365 | 6.5E-26 | 15 | 150 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 270 aa Download sequence Send to blast |
MDAAKEPAVA KPRLPPGFRF RPTDEELVVH YLRRRALASP LPAAVDIPDV RILAHDPSDL 60 LPPGWSEQER YFFTCKEAKY VKGRRANRAT GAGYWKATGK EMPVAVAVPA RGGGGQAQAV 120 LVGMKRSLVF YRGKPPTGSK TDWVMHEYRL AGAGLGPCRG AAARPAEGWV LCRVFRKKGS 180 ASANAAAAAA EQGSSGDVEE AEEAEEEDGA AGGARTFIDF FARADADAAG RREEQGQRRA 240 SSPVVSSSCL TDASHEQHGR EQETTSRGA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 5e-38 | 14 | 181 | 17 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 5e-38 | 14 | 181 | 17 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 5e-38 | 14 | 181 | 17 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 5e-38 | 14 | 181 | 17 | 169 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 6e-38 | 14 | 181 | 20 | 172 | NAC domain-containing protein 19 |
| 3swm_B | 6e-38 | 14 | 181 | 20 | 172 | NAC domain-containing protein 19 |
| 3swm_C | 6e-38 | 14 | 181 | 20 | 172 | NAC domain-containing protein 19 |
| 3swm_D | 6e-38 | 14 | 181 | 20 | 172 | NAC domain-containing protein 19 |
| 3swp_A | 6e-38 | 14 | 181 | 20 | 172 | NAC domain-containing protein 19 |
| 3swp_B | 6e-38 | 14 | 181 | 20 | 172 | NAC domain-containing protein 19 |
| 3swp_C | 6e-38 | 14 | 181 | 20 | 172 | NAC domain-containing protein 19 |
| 3swp_D | 6e-38 | 14 | 181 | 20 | 172 | NAC domain-containing protein 19 |
| 4dul_A | 5e-38 | 14 | 181 | 17 | 169 | NAC domain-containing protein 19 |
| 4dul_B | 5e-38 | 14 | 181 | 17 | 169 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.20702 | 0.0 | root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Up-regulated during xylem vessel element differentiation. {ECO:0000269|PubMed:20388856}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in xylem and phloem cells in roots and inflorescence stems (PubMed:20388856). Highly expressed in senescent leaves. Expressed in roots, and abscission and dehiscence tissues, such as axils of bracts and abscission zones in cauline leaves and siliques (PubMed:21673078). {ECO:0000269|PubMed:20388856, ECO:0000269|PubMed:21673078}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor that negatively regulates the expression of genes involved in xylem vessel formation. Represses the transcriptional activation activity of NAC030/VND7, which regulates protoxylem vessel differentiation by promoting immature xylem vessel-specific genes expression (PubMed:20388856). Transcriptional activator that regulates the COLD-REGULATED (COR15A and COR15B) and RESPONSIVE TO DEHYDRATION (LTI78/RD29A and LTI65/RD29B) genes by binding directly to their promoters. Mediates signaling crosstalk between salt stress response and leaf aging process (PubMed:21673078). May play a role in DNA replication of mungbean yellow mosaic virus (PubMed:24442717). {ECO:0000269|PubMed:20388856, ECO:0000269|PubMed:21673078, ECO:0000269|PubMed:24442717}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00507 | DAP | Transfer from AT5G13180 | Download |
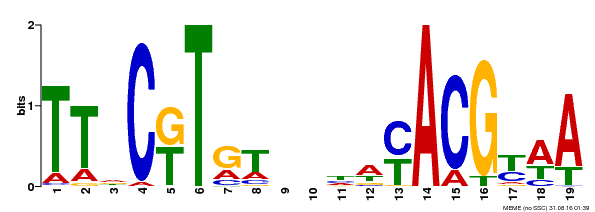
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.3NG212900.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA) and salt stress. {ECO:0000269|PubMed:21673078}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025809222.1 | 1e-139 | NAC domain-containing protein 83 | ||||
| Swissprot | Q9FY93 | 1e-56 | NAC83_ARATH; NAC domain-containing protein 83 | ||||
| TrEMBL | A0A3L6RAA2 | 1e-149 | A0A3L6RAA2_PANMI; NAC domain-containing protein 83 | ||||
| STRING | Pavir.Cb00769.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6317 | 32 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G13180.1 | 3e-44 | NAC domain containing protein 83 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.3NG212900.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



