 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G13180.1 | ||||||||
| Common Name | ANAC083, NAC083, T19L5.140, VNI2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 252aa MW: 28569.9 Da PI: 9.1386 | ||||||||
| Description | NAC domain containing protein 83 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 165.4 | 2e-51 | 14 | 138 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlsk.kg 97
lppGfrFhPtdeelvv+yLk+kv +++l++ ++i+e+d+++++PwdLp + eke yfFs+r+ ky++g+r+nrat sgyWkatg dk+v+++ ++
AT5G13180.1 14 LPPGFRFHPTDEELVVQYLKRKVCSSPLPA-SIIPEFDVCRADPWDLPGN---LEKERYFFSTREAKYPNGNRSNRATGSGYWKATGIDKRVVTSrGN 107
79****************************.89***************44...4789********************************999988577 PP
NAM 98 elvglkktLvfykgrapkgektdWvmheyrl 128
+ vglkktLvfykg+ p+g++tdW+mheyrl
AT5G13180.1 108 QIVGLKKTLVFYKGKPPHGSRTDWIMHEYRL 138
77***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 2.75E-60 | 10 | 160 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 58.558 | 14 | 160 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.0E-27 | 15 | 138 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0016032 | Biological Process | viral process | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000272 | anatomy | protoxylem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0005421 | anatomy | parenchyma | ||||
| PO:0006077 | anatomy | protophloem | ||||
| PO:0006307 | anatomy | root procambium | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0009053 | anatomy | peduncle | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 252 aa Download sequence Send to blast |
MDNVKLVKNG VLRLPPGFRF HPTDEELVVQ YLKRKVCSSP LPASIIPEFD VCRADPWDLP 60 GNLEKERYFF STREAKYPNG NRSNRATGSG YWKATGIDKR VVTSRGNQIV GLKKTLVFYK 120 GKPPHGSRTD WIMHEYRLSS SPPSSMGPTQ NWVLCRIFLK KRAGNKNDDD DGDSRNLRHN 180 NNNNSSDQIE IITTDQTDDK TKPIFFDFMR KERTTDLNLL PSSPSSDHAS SGVTTEIFSS 240 SDEETSSCNS FR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 1e-52 | 12 | 166 | 15 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 1e-52 | 12 | 166 | 15 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 1e-52 | 12 | 166 | 15 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 1e-52 | 12 | 166 | 15 | 171 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 1e-52 | 12 | 166 | 18 | 174 | NAC domain-containing protein 19 |
| 3swm_B | 1e-52 | 12 | 166 | 18 | 174 | NAC domain-containing protein 19 |
| 3swm_C | 1e-52 | 12 | 166 | 18 | 174 | NAC domain-containing protein 19 |
| 3swm_D | 1e-52 | 12 | 166 | 18 | 174 | NAC domain-containing protein 19 |
| 3swp_A | 1e-52 | 12 | 166 | 18 | 174 | NAC domain-containing protein 19 |
| 3swp_B | 1e-52 | 12 | 166 | 18 | 174 | NAC domain-containing protein 19 |
| 3swp_C | 1e-52 | 12 | 166 | 18 | 174 | NAC domain-containing protein 19 |
| 3swp_D | 1e-52 | 12 | 166 | 18 | 174 | NAC domain-containing protein 19 |
| 4dul_A | 1e-52 | 12 | 166 | 15 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 1e-52 | 12 | 166 | 15 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.10079 | 0.0 | root | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 145357958 | 0.0 | ||||
| Genevisible | 245987_at | 0.0 | ||||
| Expression Atlas | AT5G13180 | - | ||||
| AtGenExpress | AT5G13180 | - | ||||
| ATTED-II | AT5G13180 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Up-regulated during xylem vessel element differentiation. {ECO:0000269|PubMed:20388856}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in xylem and phloem cells in roots and inflorescence stems (PubMed:20388856). Highly expressed in senescent leaves. Expressed in roots, and abscission and dehiscence tissues, such as axils of bracts and abscission zones in cauline leaves and siliques (PubMed:21673078). {ECO:0000269|PubMed:20388856, ECO:0000269|PubMed:21673078}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a NAC domain transcription factor that interacts with VND7 and negatively regulates xylem vessel formation. | |||||
| UniProt | Transcriptional repressor that negatively regulates the expression of genes involved in xylem vessel formation. Represses the transcriptional activation activity of NAC030/VND7, which regulates protoxylem vessel differentiation by promoting immature xylem vessel-specific genes expression (PubMed:20388856). Transcriptional activator that regulates the COLD-REGULATED (COR15A and COR15B) and RESPONSIVE TO DEHYDRATION (LTI78/RD29A and LTI65/RD29B) genes by binding directly to their promoters. Mediates signaling crosstalk between salt stress response and leaf aging process (PubMed:21673078). May play a role in DNA replication of mungbean yellow mosaic virus (PubMed:24442717). {ECO:0000269|PubMed:20388856, ECO:0000269|PubMed:21673078, ECO:0000269|PubMed:24442717}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
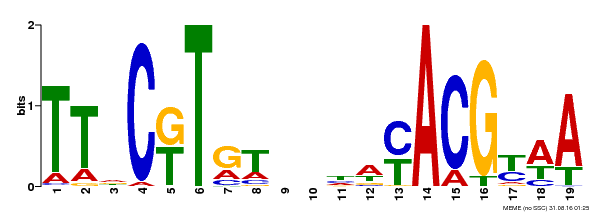
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00507 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G13180.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA) and salt stress. {ECO:0000269|PubMed:21673078}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| BioGRID | AT5G53950, AT5G62380, AT5G63790, AT5G66300, AT1G56010, AT1G71930 | |||||
| IntAct | Search Q9FY93 | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: Accelerated leaf aging. {ECO:0000269|PubMed:21673078}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G13180 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF385734 | 0.0 | AF385734.1 Arabidopsis thaliana AT5g13180/T19L5_140 mRNA, complete cds. | |||
| GenBank | AY143964 | 0.0 | AY143964.1 Arabidopsis thaliana At5g13180/T19L5_140 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_196822.1 | 0.0 | NAC domain containing protein 83 | ||||
| Swissprot | Q9FY93 | 0.0 | NAC83_ARATH; NAC domain-containing protein 83 | ||||
| TrEMBL | A0A178UJZ2 | 0.0 | A0A178UJZ2_ARATH; VNI2 | ||||
| STRING | AT5G13180.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1806 | 27 | 86 | Representative plant | OGRP17 | 15 | 800 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G13180.1 |
| Entrez Gene | 831157 |
| iHOP | AT5G13180 |
| wikigenes | AT5G13180 |



