 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.2NG054200.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | CPP | ||||||||
| Protein Properties | Length: 777aa MW: 83652.4 Da PI: 6.582 | ||||||||
| Description | CPP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCR | 50.2 | 4.9e-16 | 462 | 500 | 2 | 40 |
TCR 2 ekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
+k+C+CkkskClk+YCeCfaag++Cse C+C++C Nk
Pavir.2NG054200.1.p 462 ACKRCSCKKSKCLKLYCECFAAGVYCSEPCSCQGCLNKP 500
589**********************************96 PP
| |||||||
| 2 | TCR | 50.2 | 5e-16 | 547 | 586 | 1 | 40 |
TCR 1 kekkgCnCkkskClkkYCeCfaagkkCseeCkCedCkNke 40
++k+gCnCkks+ClkkYCeC++ g+ Cs++C+Ce+CkN+
Pavir.2NG054200.1.p 547 RHKRGCNCKKSSCLKKYCECYQGGVGCSSNCRCESCKNTF 586
589***********************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01114 | 3.8E-17 | 461 | 502 | IPR033467 | Tesmin/TSO1-like CXC domain |
| PROSITE profile | PS51634 | 37.046 | 462 | 587 | IPR005172 | CRC domain |
| Pfam | PF03638 | 8.8E-12 | 464 | 499 | IPR005172 | CRC domain |
| SMART | SM01114 | 3.5E-16 | 547 | 588 | IPR033467 | Tesmin/TSO1-like CXC domain |
| Pfam | PF03638 | 1.1E-11 | 549 | 585 | IPR005172 | CRC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009934 | Biological Process | regulation of meristem structural organization | ||||
| GO:0048444 | Biological Process | floral organ morphogenesis | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 777 aa Download sequence Send to blast |
MDTPDRAAAQ PPPASRAEDS PVFSFINNLS PIEPLKSAYN ASSIQGYQSI NITSVSSIFT 60 SPHDNAHKET RLAKSSLGEI SESEVCAGGS KTNKPAKSSN AVRLFARAST VTRETHTVTC 120 SDVADPPTEP CDLAQPAQFD NGSPDHNTTP CHGVRSDLKQ EKCRKLDVVQ TVKNTVEKRK 180 CLFSTEIQLL DGGQPVNDSD DVLGCEWSDL ISTTSGELLA FDSTMDDHHR GMHLAAKNAE 240 SCGYLLSKLT GGGEISDRAH PSASGQVYYQ EIVMGEDQAE NPQIFQDGQQ TISTEEIQDN 300 IYEANGCIPL DYKVESQQRG IRRRCLVFEA AGFSNTVVQK ETVESLSVST CKGKSHVQTQ 360 PRGLRGIGLH LNALALTPKG KMASQDPMAS GLHPSSGSEK DVHGKFLSAG EKFPDSGGEL 420 LEFPMDDCSA GGFPVNDHVS SQSVSPQKKR RKTDNNGDDG EACKRCSCKK SKCLKLYCEC 480 FAAGVYCSEP CSCQGCLNKP IHEEIVLSTR KQIEFRNPLA FAPKVIRMSD AGPETGEDPN 540 STPASARHKR GCNCKKSSCL KKYCECYQGG VGCSSNCRCE SCKNTFGRRD ADTELTEEMK 600 QEGEQAENCG KEKENDQQKA NLQNEDHPLL ELVPITPPFD LSSSLLKLPN FSSAKPPRPS 660 KARSGNSRSS ASKVTATLQS CKSSKVAGSA IDEEMPDILK EADSPNNCVK TTSPNGKRVS 720 PPHNALSISP NRKGGRKLIL KSIPSFPSLM GESNSGSTMN DTENAFSTSP LVLGPS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5fd3_A | 3e-15 | 464 | 584 | 12 | 120 | Protein lin-54 homolog |
| 5fd3_B | 3e-15 | 464 | 584 | 12 | 120 | Protein lin-54 homolog |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 447 | 451 | KKRRK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.13912 | 0.0 | leaf| stem | ||||
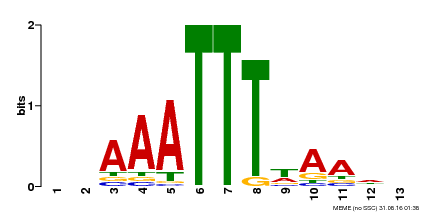
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00624 | PBM | Transfer from PK22848.1 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.2NG054200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025799594.1 | 0.0 | protein tesmin/TSO1-like CXC 2 | ||||
| TrEMBL | A0A2S3GWC5 | 0.0 | A0A2S3GWC5_9POAL; Uncharacterized protein | ||||
| TrEMBL | A0A2T7EL85 | 0.0 | A0A2T7EL85_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Bb00411.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6147 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22780.1 | 2e-88 | Tesmin/TSO1-like CXC domain-containing protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.2NG054200.1.p |



