 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pavir.1NG010700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 865aa MW: 97403.9 Da PI: 7.8337 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13 | 0.0003 | 558 | 580 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C Cg +F + +Lk+H+++H
Pavir.1NG010700.1.p 558 HRCLECGACFQKPAHLKQHMQSH 580
79*******************99 PP
| |||||||
| 2 | zf-C2H2 | 22.4 | 3.1e-07 | 586 | 610 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s++rk++L+rH+ +H
Pavir.1NG010700.1.p 586 FICPleDCPFSYKRKDHLNRHMLKH 610
78*********************99 PP
| |||||||
| 3 | zf-C2H2 | 11.3 | 0.0011 | 617 | 640 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirt.H 23
C+ C+++Fs k n++rH + H
Pavir.1NG010700.1.p 617 CTvdGCDRRFSIKANMQRHVKEfH 640
88889***************9988 PP
| |||||||
| 4 | zf-C2H2 | 15.2 | 6.1e-05 | 652 | 676 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ C+ C+k F+ s Lk+H +H
Pavir.1NG010700.1.p 652 FICKeeGCNKAFKYLSKLKKHEESH 676
789999***************9988 PP
| |||||||
| 5 | zf-C2H2 | 17.7 | 1e-05 | 743 | 767 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C+ sFs+ksnL++H++ H
Pavir.1NG010700.1.p 743 KCTfdGCDHSFSSKSNLTKHMKAcH 767
799999****************955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.25.40.10 | 1.7E-7 | 27 | 164 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 5.02 | 35 | 65 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.388 | 66 | 100 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 1 | 68 | 97 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.046 | 137 | 167 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.0021 | 142 | 164 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 9.745 | 168 | 202 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.018 | 170 | 198 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 2.1E-4 | 201 | 234 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 4.2E-4 | 201 | 230 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 6.752 | 203 | 233 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 1.7E-7 | 256 | 380 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 8.55 | 269 | 299 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 0.0016 | 274 | 299 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF13041 | 4.5E-11 | 299 | 347 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 11.959 | 300 | 334 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 7.9E-9 | 302 | 335 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 9.098 | 335 | 365 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 0.0027 | 337 | 365 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 6.621 | 371 | 401 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.018 | 374 | 398 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 1.7E-7 | 435 | 465 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 7.015 | 437 | 471 | IPR002885 | Pentatricopeptide repeat |
| SMART | SM00355 | 19 | 512 | 534 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.9E-4 | 555 | 577 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 12.591 | 558 | 585 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.04 | 558 | 580 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 560 | 580 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.78E-10 | 570 | 612 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-11 | 578 | 612 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.037 | 586 | 615 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.9E-4 | 586 | 610 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 588 | 610 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.51E-7 | 596 | 646 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.7E-8 | 613 | 642 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.512 | 615 | 645 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.027 | 615 | 640 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 617 | 640 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-6 | 649 | 676 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.201 | 652 | 676 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0087 | 652 | 676 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 654 | 676 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 6.8E-4 | 684 | 704 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.7 | 684 | 709 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 686 | 709 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 13 | 712 | 733 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.5E-6 | 727 | 764 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 6.15E-9 | 727 | 784 | No hit | No description |
| PROSITE profile | PS50157 | 11.863 | 742 | 772 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.8E-4 | 742 | 767 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 744 | 767 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.1E-5 | 771 | 795 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.245 | 773 | 799 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 2.6 | 773 | 799 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 775 | 799 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 865 aa Download sequence Send to blast |
MNRSICHHLL SQCKTLRELQ KIHAQAVAQG LHPQHQPVSC KIFRCYADFG RAADARKLFD 60 EIPRPDLVSF TSLMSLHIRL EHHRAAVSLF SRVVADGHHR PDGFAVVGAL SASGGAGDQQ 120 VGRAVHGMIF RLGLDYEVVV GNALVDMYSR CGKFESALLV FDRMFLKDEI TWGSMLHGYI 180 KCAGLDLALS FFDQMPVKSV VAWTALITGH VQGRLPVRAL ELFGRMVLEG HRPTHVTVVG 240 VFSACADIGA LDLGRIIHGY GTKCNASSNI IVSNALMDMY AKSGSIEMAF SVFQEVQSKD 300 AYTWTTMISC FTVQGDGRKA LELFRDMLRS SVVPNSVTFV SVLSACSHAG LIEEGRELFG 360 TMREMYSIDP QLEHYGCMID LLGRGGLLEE AEALIVDMNV EPDIVIWRSL LSASLVHRND 420 RLAEIAGKEI MKREPGDDGV YVLLWNMYAS SNKWIEAREI RQQMLTRKIF KKPGCSWIEV 480 DGVVHEFLMC SGDEIDGDAS VGQKGFKDIR CYKCEFCTIV RSKKCLIQAH MIAHHKDELE 540 KSETYNSNGE KIVCEEEHRC LECGACFQKP AHLKQHMQSH SNERLFICPL EDCPFSYKRK 600 DHLNRHMLKH QGKLFGCTVD GCDRRFSIKA NMQRHVKEFH EDENVTKSNQ QFICKEEGCN 660 KAFKYLSKLK KHEESHVKLN YTEVVCCEPG CMKMFTNVES LRAHNQSCHQ HVQCEICGKK 720 HLKKNIKRHL QAHDELASGE RIKCTFDGCD HSFSSKSNLT KHMKACHDQL KPFTCRVAGC 780 GKAFTYKHVR DNHEKSSAHV YVEGDFVETD EQLRLRPRGG RKRKALSVET LTRKRVTILG 840 EASSLEDGAE YLSWLLSGGD DSAQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5iww_D | 3e-53 | 9 | 413 | 3 | 338 | PLS9-PPR |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 815 | 823 | RPRGGRKRK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pvr.19738 | 0.0 | callus| leaf| root| stem | ||||
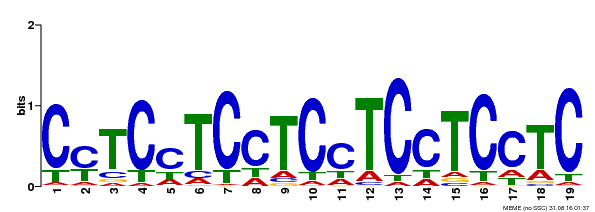
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pavir.1NG010700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025795590.1 | 0.0 | pentatricopeptide repeat-containing protein At2g22410, mitochondrial-like | ||||
| TrEMBL | K3Z0X6 | 0.0 | K3Z0X6_SETIT; Uncharacterized protein | ||||
| STRING | Pavir.J03392.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-105 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pavir.1NG010700.1.p |



