 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Potri.019G081500.1 | ||||||||
| Common Name | POPTR_0019s11090g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 299aa MW: 33730.8 Da PI: 8.7831 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.1 | 1.7e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd++l+d+++ +G g+W++ ++ g+ R++k+c++rw +yl
Potri.019G081500.1 14 KGAWTKEEDQRLIDYIRVHGEGCWRSLPKAAGLLRCGKSCRLRWINYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 59.4 | 8e-19 | 67 | 111 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg++T+eEde++++++ +lG++ W++Ia +++ gRt++++k++w+++
Potri.019G081500.1 67 RGNFTEEEDEIIIKLHSLLGNK-WSLIAGKLP-GRTDNEIKNYWNTH 111
89********************.*********.************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 7.5E-24 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.249 | 9 | 61 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.4E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 9.16E-30 | 13 | 108 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.5E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.72E-10 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 29.286 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-28 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 7.8E-18 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-16 | 67 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.28E-12 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
MGRSPCCEKA HTNKGAWTKE EDQRLIDYIR VHGEGCWRSL PKAAGLLRCG KSCRLRWINY 60 LRPDLKRGNF TEEEDEIIIK LHSLLGNKWS LIAGKLPGRT DNEIKNYWNT HIKRKLVSRG 120 IDPQTHRPLN EKTTTTTTAT STTKTTQLNF KNTPPQSLAE ISILKSQLDS KYNSTKSNDF 180 LSSLYSSFKP KKIETASLEE NNCTSCSTTT DEEQQQKRES HHDQDINLDL TIGLAPTTKG 240 ESTHTSSNSA ESRLQQPVSS RQLFGSVSTG PGCLCWQSVG AGWESCRDCQ NHRYYCWK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 5e-30 | 13 | 116 | 26 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. {ECO:0000269|PubMed:9839469}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00053 | PBM | Transfer from AT1G22640 | Download |
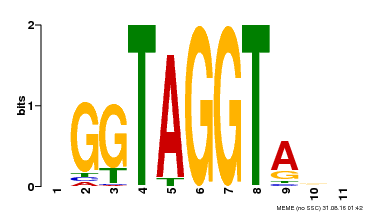
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Potri.019G081500.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By nitrogen, salicylic acid, NaCl and abscisic acid (ABA). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:18541146, ECO:0000269|PubMed:9839469}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002325546.1 | 0.0 | transcription factor MYB8 | ||||
| Swissprot | Q9S9K9 | 3e-93 | MYB3_ARATH; Transcription factor MYB3 | ||||
| TrEMBL | B9IQH2 | 0.0 | B9IQH2_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0019s11090.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF264 | 34 | 221 | Representative plant | OGRP5 | 17 | 1784 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G22640.1 | 4e-94 | myb domain protein 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Potri.019G081500.1 |
| Entrez Gene | 7475793 |



