 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Potri.012G007500.2 | ||||||||
| Common Name | POPTR_0012s00760g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 516aa MW: 57679.6 Da PI: 4.5247 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 170.4 | 5.6e-53 | 11 | 138 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp..kkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkd 89
lp GfrF+P+deelv++yL++k++gk+ ++ +vi+e+d++k+ePwdLp + +++++ ew+fF++ d+ky++g+r+nrat++gyWkatgkd
Potri.012G007500.2 11 LPLGFRFRPSDEELVNYYLRHKINGKDDKV-RVIREIDVCKWEPWDLPglSIIENKDPEWFFFCPLDRKYPNGSRQNRATNAGYWKATGKD 100
699************************999.99***************6336777888********************************* PP
NAM 90 kevlskkgelvglkktLvfykgrapkgektdWvmheyrl 128
++++s +++l+g+kktLvfy+grapkg++t+Wv+heyr+
Potri.012G007500.2 101 RKIKS-GNRLIGMKKTLVFYTGRAPKGKRTNWVVHEYRA 138
*****.999****************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 2.62E-61 | 4 | 161 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 57.1 | 11 | 161 | IPR003441 | NAC domain |
| Pfam | PF02365 | 4.3E-27 | 12 | 137 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0002230 | Biological Process | positive regulation of defense response to virus by host | ||||
| GO:0002237 | Biological Process | response to molecule of bacterial origin | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009814 | Biological Process | defense response, incompatible interaction | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0070417 | Biological Process | cellular response to cold | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0003713 | Molecular Function | transcription coactivator activity | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 516 aa Download sequence Send to blast |
MAVFSLNLNS LPLGFRFRPS DEELVNYYLR HKINGKDDKV RVIREIDVCK WEPWDLPGLS 60 IIENKDPEWF FFCPLDRKYP NGSRQNRATN AGYWKATGKD RKIKSGNRLI GMKKTLVFYT 120 GRAPKGKRTN WVVHEYRATE EELDGTKPGQ SAFVLCRLFK KQDESIESPN CDEAEATVSS 180 PTTAQSSPEV TQSDQPLTEA SPANTTTSEV VAPVEFPSCS VGVSEGGDQT ADLPASEEEL 240 QLEEALNWLF DSPPQALDYE LFPPVHEQVQ EAVGSSSMFN HGNNDLSSSN RGLQSHNVGN 300 ETDDYTSEFI DSILKQPDEL FYEVPSFQNN SSFPSESLFR GSLLRVVEDN VSYSGSDVDM 360 EPIRIEQGFQ GAVFPEGNID EKPSSTLLYD SNVHQQPIYL GSLQNGSYAL QSVASISATD 420 QLNTLNKPSN STNEVGGSDT NWNGIRTRVH NPRNRQFGRN FNTQGDAPRR LRLQCKLQVQ 480 PIQFSTKSED FNSGEGELEY ECIVTKVRSL YNDAE* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 1e-47 | 10 | 165 | 16 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 1e-47 | 10 | 165 | 16 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 1e-47 | 10 | 165 | 16 | 169 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 1e-47 | 10 | 165 | 16 | 169 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 1e-47 | 10 | 165 | 19 | 172 | NAC domain-containing protein 19 |
| 3swm_B | 1e-47 | 10 | 165 | 19 | 172 | NAC domain-containing protein 19 |
| 3swm_C | 1e-47 | 10 | 165 | 19 | 172 | NAC domain-containing protein 19 |
| 3swm_D | 1e-47 | 10 | 165 | 19 | 172 | NAC domain-containing protein 19 |
| 3swp_A | 1e-47 | 10 | 165 | 19 | 172 | NAC domain-containing protein 19 |
| 3swp_B | 1e-47 | 10 | 165 | 19 | 172 | NAC domain-containing protein 19 |
| 3swp_C | 1e-47 | 10 | 165 | 19 | 172 | NAC domain-containing protein 19 |
| 3swp_D | 1e-47 | 10 | 165 | 19 | 172 | NAC domain-containing protein 19 |
| 4dul_A | 1e-47 | 10 | 165 | 16 | 169 | NAC domain-containing protein 19 |
| 4dul_B | 1e-47 | 10 | 165 | 16 | 169 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Pth.11862 | 0.0 | bud | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator essential for the anti-viral defense called virus basal resistance response pathway (PubMed:11041886, PubMed:15629774, PubMed:18785827, PubMed:24418554). Not involved in HRT-mediated hypersensitive response (HR) and resistance to TCV (PubMed:18785827). Binds DNA non specifically (PubMed:15629774). Activated by proteolytic cleavage through regulated intramembrane proteolysis (RIP) (By similarity). {ECO:0000250|UniProtKB:Q9SCK6, ECO:0000269|PubMed:11041886, ECO:0000269|PubMed:15629774, ECO:0000269|PubMed:18785827, ECO:0000269|PubMed:24418554}.; FUNCTION: (Microbial infection) Compromised function in defense response pathway when interacting with the invading viral capsid protein (CP) of turnip crinkle virus (TCV) due to abnormal subcellular localization. {ECO:0000269|PubMed:11041886, ECO:0000269|PubMed:15629774}. | |||||
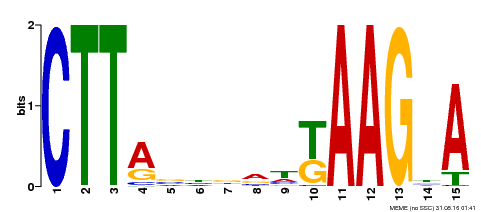
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00398 | DAP | Transfer from AT3G49530 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Potri.012G007500.2 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by bacterial pathogens type III effector proteins (TTEs). {ECO:0000269|PubMed:16553893}.; INDUCTION: (Microbial infection) Accumulates 2 days post infection with turnip crinkle virus (TCV) (PubMed:24418554). {ECO:0000269|PubMed:24418554}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002317831.3 | 0.0 | NAC domain-containing protein 91 | ||||
| Swissprot | Q9LKG8 | 1e-100 | NAC91_ARATH; NAC domain-containing protein 91 | ||||
| TrEMBL | A0A2K1Y6Z1 | 0.0 | A0A2K1Y6Z1_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0012s00760.1 | 0.0 | (Populus trichocarpa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G24590.2 | 1e-100 | TCV-interacting protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Potri.012G007500.2 |
| Entrez Gene | 7496453 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



