 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | model.Picochlorum_contig_197.g886.t1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Trebouxiophyceae incertae sedis; Picochlorum
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 534aa MW: 57701.3 Da PI: 6.3829 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 63.7 | 3.8e-20 | 182 | 230 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkr..krfslgkfgtaeeAakaaiaarkkleg 55
y+GVr+++ +g+W+AeIrdp + +r +lg+f+taeeAaka+++a+++++g
model.Picochlorum_contig_197.g886.t1 182 YRGVRQRP-WGKWAAEIRDP-----TvgARRWLGTFDTAEEAAKAYDQAARAIRG 230
9*******.**********9.....4459***********************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51032 | 23.024 | 181 | 238 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.16E-21 | 181 | 239 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 6.3E-37 | 181 | 244 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.8E-31 | 181 | 238 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.06E-29 | 182 | 238 | No hit | No description |
| Pfam | PF00847 | 1.2E-12 | 182 | 230 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.2E-11 | 182 | 193 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.2E-11 | 204 | 220 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 534 aa Download sequence Send to blast |
MRASRSKSGS NRGGTGGHNN ADLDHINVQS ASVYDTDATN LQGGRPSVVA RPGAGLPARA 60 IPSNPQEYHV PHNTRQQAQA NSNRPSVSTR SGGNHQHHYH MRSHQHRQGG NHDVSPQTAL 120 RSGGTSSVQH PTVPVSNLSS PGVYHQHGGQ GQKSSKADDG EVMVPSGVPP PVVVSKGKNK 180 TYRGVRQRPW GKWAAEIRDP TVGARRWLGT FDTAEEAAKA YDQAARAIRG NQAKCNFPLP 240 EEEEHQHHQQ KQQRMTDRRH TSERHGRSRG REEQEEDVVA ASSISFLNPS SVPEEALIHN 300 PLAAIHKTSG VICMSEAMCI PGGSTGDEGP LVEAAADQSD MEPWEAAREP TGLTPKGGGW 360 MGPEWMPKSM NMSGGANVGS MPFGRSVEMS ADLPQKFLAS GYDTFTDMGS IRQTLEIPAD 420 FVEESDPDDD LDDDVMILGT TPQFGSTPRH PHSVLGGNKE FKHGSDKVEP GEKKTRMTTR 480 MRAMAVVVQE ETEDDDYDDS SDDNSCMLGM SPDVAEHGFS SFGAQHAEVS ARR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 1e-20 | 180 | 239 | 1 | 61 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00623 | PBM | Transfer from PK19363.1 | Download |
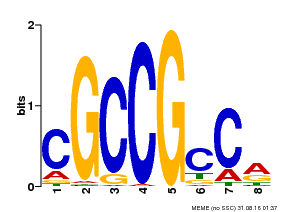
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP132 | 15 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33710.1 | 8e-19 | ERF family protein | ||||



