 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Prupe.7G018400.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 363aa MW: 41363.5 Da PI: 7.2981 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.2 | 1.7e-17 | 49 | 96 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++++G+g+W++++ g+ R+ k+c++rw +yl
Prupe.7G018400.1.p 49 KGPWTPEEDIILVSYIQEHGPGNWRSVPTNTGLLRCSKSCRLRWTNYL 96
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 45.5 | 1.7e-14 | 102 | 147 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T+ E+ ++++ ++lG++ W++Ia++++ Rt++++k++w+++l
Prupe.7G018400.1.p 102 RGNFTPHEEGMIIHLQALLGNK-WAAIASYLP-QRTDNDIKNYWNTHL 147
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-23 | 40 | 99 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 25.075 | 44 | 100 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.47E-30 | 46 | 143 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.4E-14 | 48 | 98 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.7E-16 | 49 | 96 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.09E-11 | 51 | 96 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 8.3E-25 | 100 | 152 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.75 | 101 | 151 | IPR017930 | Myb domain |
| SMART | SM00717 | 3.5E-13 | 101 | 149 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 6.5E-13 | 102 | 147 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.03E-9 | 104 | 147 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 363 aa Download sequence Send to blast |
MLITLFEIKC EVILDWKRRQ KRERERERER ERERDMGRPP CCEKVGIKKG PWTPEEDIIL 60 VSYIQEHGPG NWRSVPTNTG LLRCSKSCRL RWTNYLRPGI KRGNFTPHEE GMIIHLQALL 120 GNKWAAIASY LPQRTDNDIK NYWNTHLKKK LKKFQSALDP HMASDSTTST NHQLNIVSRR 180 SLSEGSLDNM ATHASSVTLK NNQSSTYASS TANISRLLEG WMRSSPTPKM LSNTNHLKLN 240 ASHDLQDQET PAGNLLSRHE ELESILSYEN LNNVASWDRS TCDSTQSDNK LNPRCGRVAN 300 DDEIIHEDDL VPDRDQKNKQ IIRCEGSTSA PPLSFLEKWL LDESAGQVDD DQMLELSPMS 360 SV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 2e-23 | 47 | 151 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 23 | 33 | RERERERERER |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: During seed development, gradually down-regulated towards the onset of ripening (veraison). During berry skin development, dramatic decrease to full repression at veraison, followed by a slight increase towards ripening. In flowers, barely detectable in stamens, at the interface of filaments and anthers. {ECO:0000269|PubMed:22018045}. | |||||
| Uniprot | TISSUE SPECIFICITY: Restricted to stomatal guard cells. Mostly expressed in leaves, cotyledons, hypocotyls, seeds and ripened berry skins. {ECO:0000269|PubMed:22018045}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the regulation of gene (e.g. drought-regulated and flavonoid biosynthetic genes) expression and stomatal movements leading to negative regulation of responses to drought and responses to other physiological stimuli (e.g. light). {ECO:0000269|PubMed:22018045}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
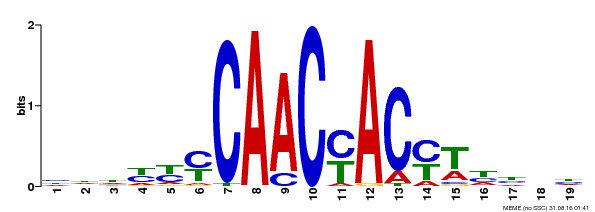
| Motif ID | Method | Source | Motif file |
| MP00132 | DAP | Transfer from AT1G08810 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Prupe.7G018400.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by abscisic acid (ABA) and osmotic stress (salt stress). {ECO:0000269|PubMed:22018045}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KP229564 | 1e-116 | KP229564.1 UNVERIFIED: Boehmeria nivea MYB transcription factor-like (MYB08) mRNA, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007203968.2 | 0.0 | myb-related protein 306 | ||||
| Swissprot | B3VTV7 | 1e-125 | MYB60_VITVI; Transcription factor MYB60 | ||||
| TrEMBL | A0A251N540 | 0.0 | A0A251N540_PRUPE; Uncharacterized protein | ||||
| STRING | EMJ05167 | 0.0 | (Prunus persica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8203 | 31 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G08810.1 | 1e-106 | myb domain protein 60 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Prupe.7G018400.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



