 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pp3c12_24350V3.1.p | ||||||||
| Common Name | PHYPADRAFT_83876 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Bryophytina; Bryopsida; Funariidae; Funariales; Funariaceae; Physcomitrella
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 687aa MW: 73539.4 Da PI: 7.4171 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.8 | 1.2e-41 | 217 | 293 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cq+egC+ dls+ak+yhrrhkvCe hskap+v+v g++qrfCqqCsrfh+l efDe+krsCr+rLa+hn+rrrk+q
Pp3c12_24350V3.1.p 217 VCQAEGCKDDLSNAKHYHRRHKVCELHSKAPTVTVGGHTQRFCQQCSRFHHLGEFDEGKRSCRKRLADHNRRRRKPQ 293
6**************************************************************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 5.6E-35 | 211 | 279 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.436 | 215 | 292 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 6.67E-39 | 216 | 295 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.2E-31 | 218 | 291 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0025017 | anatomy | plant spore | ||||
| PO:0030003 | anatomy | protonema | ||||
| PO:0030018 | anatomy | gametophore | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 687 aa Download sequence Send to blast |
MNPSASSNEQ QGDPSWATEN WDNPGAVGSL DQASRQYTGS NRQEWEWDPM ILAQHPGSGG 60 SNDGSDGDRK NQISAGAMSS LRASYNMLSA GLPSNFTHNP LGIFSSAGIR NFGSLDGSGQ 120 ISTSANLHSV LGSSGIPGLA ATSGTGFCND IRSYERRCDF FAANMHDHRV KREEVSDGHA 180 RIGLNLGVRT YFSTEDTAVG RLGKRHRAGS PGFQVPVCQA EGCKDDLSNA KHYHRRHKVC 240 ELHSKAPTVT VGGHTQRFCQ QCSRFHHLGE FDEGKRSCRK RLADHNRRRR KPQLNASTSG 300 GTSVESIGII KACDDGHLHA GINRPNDPKS MLNMHSQKNC GSPNSVSLED SDEKPSLGAK 360 NLKLRTSGPG YCRSPGNDQQ SSHSGMQMDS QSMASSEAQT LLSAISLSPP APLVQHKSKR 420 QLTMLGGTGD YDHENSVYQL YLQGVSSSQT GPNLSLSSLE SQLGHQGGGR GQPTSVQSYN 480 GMESAVQWLR PINAQSDSMT KAVNRSGAMN MQHLISTDSK SGITSASANS SNSQEHDVQA 540 FSPSNQSLLP LESRDLSSPE WMMGSITGAQ TSRDGSHQYR PSTPMNNLSA SEQLEAGDRM 600 LALINTSNQA GSNPGNDGSN QNVLTGRSQM EFLQQQNTGA DSGGDNNSRG GGSTVNSPDL 660 KYPELQSLRP SSYGTSSIYD SHHNLL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-29 | 208 | 291 | 3 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 274 | 291 | KRSCRKRLADHNRRRRKP |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
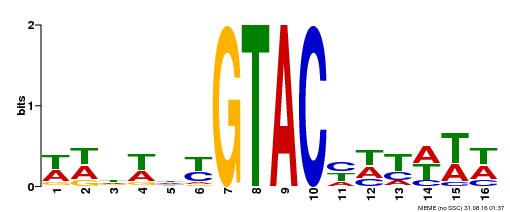
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ968320 | 0.0 | AJ968320.1 Physcomitrella patens SBP1 gene for squamosa promoter binding protein 1. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024390961.1 | 0.0 | squamosa promoter-binding-like protein 14 | ||||
| Refseq | XP_024390962.1 | 0.0 | squamosa promoter-binding-like protein 14 | ||||
| Refseq | XP_024390963.1 | 0.0 | squamosa promoter-binding-like protein 14 | ||||
| Refseq | XP_024390964.1 | 0.0 | squamosa promoter-binding-like protein 14 | ||||
| Refseq | XP_024390965.1 | 0.0 | squamosa promoter-binding-like protein 14 | ||||
| Refseq | XP_024390966.1 | 0.0 | squamosa promoter-binding-like protein 14 | ||||
| Refseq | XP_024390967.1 | 0.0 | squamosa promoter-binding-like protein 14 | ||||
| Refseq | XP_024390968.1 | 0.0 | squamosa promoter-binding-like protein 14 | ||||
| TrEMBL | A9SU35 | 0.0 | A9SU35_PHYPA; Predicted protein | ||||
| STRING | PP1S118_9V6.1 | 0.0 | (Physcomitrella patens) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP97 | 17 | 230 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 4e-49 | squamosa promoter binding protein-like 8 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pp3c12_24350V3.1.p |
| Entrez Gene | 5933082 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



