 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pp3c12_14300V3.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Bryophyta; Bryophytina; Bryopsida; Funariidae; Funariales; Funariaceae; Physcomitrella
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 357aa MW: 38310.3 Da PI: 10.1403 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 44.5 | 3.6e-14 | 89 | 133 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++lG+g+W+ I+r + ++Rt+ q+ s+ qky
Pp3c12_14300V3.1.p 89 PWTEEEHRLFLLGLQKLGKGDWRGISRNFVQTRTPTQVASHAQKY 133
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.60.10 | 9.2E-4 | 4 | 21 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS50158 | 9.026 | 5 | 20 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS51294 | 19.91 | 82 | 138 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.72E-18 | 84 | 139 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.9E-18 | 85 | 137 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-11 | 86 | 132 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.6E-11 | 86 | 136 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.06E-10 | 89 | 134 | No hit | No description |
| Pfam | PF00249 | 1.2E-11 | 89 | 133 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000122 | Biological Process | negative regulation of transcription from RNA polymerase II promoter | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0030307 | Biological Process | positive regulation of cell growth | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:2000469 | Biological Process | negative regulation of peroxidase activity | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 357 aa Download sequence Send to blast |
MARGCSHCGH NGHNSRTCPD RGVRLFGVRL TDGVMRKSVS MGNLSHYASP NNPSSPPSHS 60 ESGAGGDGYV SDGLVQTSNN TRERKKGVPW TEEEHRLFLL GLQKLGKGDW RGISRNFVQT 120 RTPTQVASHA QKYFIRQSNI NKRKRRSSLF DIVSETGPTP ILEEPTTKAV PDMSAPLHQL 180 SLGPNSTYPG VFYDPNSSHG FIRPFMLPPT SLAVPIMSMG PVALGASEQI PETSAVNFNG 240 DAARAMRPMG IPVSGPSGAM GIPYPFPMFS MLPRGYNRPV NSADSKVLRP TAKLSTEPLN 300 VGVDETKDMS QLNLGLSTPE PSQLTLKLLD QPSRSSSAFH VSTSSINSSS NAIGVV* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ppa.8236 | 0.0 | gametophore| protonema | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
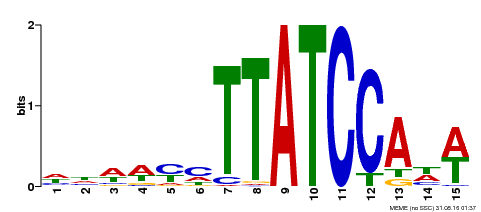
| Motif ID | Method | Source | Motif file |
| MP00551 | DAP | Transfer from AT5G47390 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024390573.1 | 0.0 | transcription factor KUA1-like isoform X1 | ||||
| Refseq | XP_024390574.1 | 0.0 | transcription factor KUA1-like isoform X1 | ||||
| TrEMBL | A0A2K1JQS5 | 0.0 | A0A2K1JQS5_PHYPA; Uncharacterized protein | ||||
| STRING | PP1S70_203V6.1 | 0.0 | (Physcomitrella patens) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP394 | 16 | 99 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47390.1 | 8e-67 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pp3c12_14300V3.1.p |



