 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G47390.1 | ||||||||
| Common Name | MYBH | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 365aa MW: 39670.9 Da PI: 6.5656 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 41.6 | 3e-13 | 97 | 141 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ ++ + ++lG+g+W+ I+r ++Rt+ q+ s+ qky
AT5G47390.1 97 PWTEEEHRMFLLGLQKLGKGDWRGISRNYVTTRTPTQVASHAQKY 141
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50158 | 8.515 | 3 | 20 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS51294 | 19.44 | 90 | 146 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.01E-17 | 92 | 147 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.8E-18 | 93 | 145 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.6E-11 | 94 | 144 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-11 | 94 | 140 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.70E-9 | 97 | 142 | No hit | No description |
| Pfam | PF00249 | 7.0E-11 | 97 | 141 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000122 | Biological Process | negative regulation of transcription from RNA polymerase II promoter | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0030307 | Biological Process | positive regulation of cell growth | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:2000469 | Biological Process | negative regulation of peroxidase activity | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000084 | anatomy | plant sperm cell | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025195 | anatomy | pollen tube cell | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001016 | developmental stage | L mature pollen stage | ||||
| PO:0001017 | developmental stage | M germinated pollen stage | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 365 aa Download sequence Send to blast |
MTRRCSHCNH NGHNSRTCPN RGVKLFGVRL TEGSIRKSAS MGNLSHYTGS GSGGHGTGSN 60 TPGSPGDVPD HVAGDGYASE DFVAGSSSSR ERKKGTPWTE EEHRMFLLGL QKLGKGDWRG 120 ISRNYVTTRT PTQVASHAQK YFIRQSNVSR RKRRSSLFDM VPDEVGDIPM DLQEPEEDNI 180 PVETEMQGAD SIHQTLAPSS LHAPSILEIE ECESMDSTNS TTGEPTATAA AASSSSRLEE 240 TTQLQSQLQP QPQLPGSFPI LYPTYFSPYY PFPFPIWPAG YVPEPPKKEE THEILRPTAV 300 HSKAPINVDE LLGMSKLSLA ESNKHGESDQ SLSLKLGGGS SSRQSAFHPN PSSDSSDIKS 360 VIHAL |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.23392 | 0.0 | floral meristem| flower| leaf| root| seed | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 42568369 | 0.0 | ||||
| Genevisible | 248795_at | 0.0 | ||||
| Expression Atlas | AT5G47390 | - | ||||
| AtGenExpress | AT5G47390 | - | ||||
| ATTED-II | AT5G47390 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Accumulates during leaf expansion. First observed at the tip of the leaves 12 days after sowing (DAS). At 14 DAS, expressed throughout the leaf blade to fade out thereafter in a basipetal manner. In mature leaves, detected in vascular tissue, especially in companion cells (PubMed:24806884). Accumulates to higher levels in old rosette leaves than in young rosette and cauline leaves (PubMed:25920996). {ECO:0000269|PubMed:24806884, ECO:0000269|PubMed:25920996}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed ubiquitously, except in hypocotyls, root tips and lateral root primordia. {ECO:0000269|PubMed:25920996}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor (PubMed:23888064, PubMed:24806884). Direct regulator of the transcription of peroxidase (Prxs) and reactive oxygen species (ROS)-related genes via the recognition of 5'-ATCACA-3' motif (PubMed:24806884). Binds to 5'-TATCCA-3' motif (TA box) and represses the activity of corresponding promoters (e.g. sugar response genes) (PubMed:25920996). Regulates hypocotyl elongation in response to darkness by enhancing auxin accumulation in a phytochrome-interacting factor (PIF) proteins-dependent manner. Promotes lateral roots formation (PubMed:23888064). Promotes cell expansion during leaves development via the modulation of cell wall-located Prxs (PubMed:24806884). Plays a critical role in developmentally regulated and dark-induced onset of leaf senescence by repressing the transcription of several genes involved in chloroplast function and responses to light and auxin. Promotes responses to auxin, abscisic acid (ABA), and ethylene (PubMed:25920996). {ECO:0000269|PubMed:23888064, ECO:0000269|PubMed:24806884, ECO:0000269|PubMed:25920996}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00551 | DAP | 27203113 | Download |
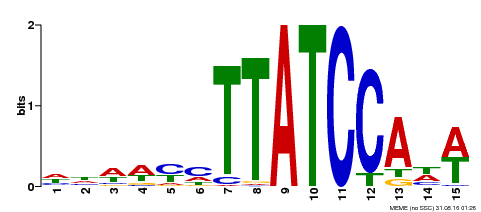
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G47390.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Slightly induced by CdCl(2) (PubMed:16463103). Accumulates in the dark (PubMed:23888064, PubMed:25920996). Diurnal expression pattern with maximal levels in the morning (at protein level). Specifically induced during leaf expansion (PubMed:24806884). Expressed in old and dark-treated leaves (PubMed:25920996). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:23888064, ECO:0000269|PubMed:24806884, ECO:0000269|PubMed:25920996}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | abscisic acid, ethylene, gibberellin, jasmonic acid, salicylic acid | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G47390 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK226931 | 0.0 | AK226931.1 Arabidopsis thaliana mRNA for Myb-related transcription activator-like, complete cds, clone: RAFL09-26-D04. | |||
| GenBank | AY072077 | 0.0 | AY072077.1 Arabidopsis thaliana putative Myb-related transcription activator protein (At5g47390) mRNA, complete cds. | |||
| GenBank | AY084295 | 0.0 | AY084295.1 Arabidopsis thaliana clone 102806 mRNA, complete sequence. | |||
| GenBank | AY096661 | 0.0 | AY096661.1 Arabidopsis thaliana putative Myb-related transcription activator (At5g47390) mRNA, complete cds. | |||
| GenBank | AY519516 | 0.0 | AY519516.1 Arabidopsis thaliana MYB transcription factor (At5g47390) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_199550.1 | 0.0 | myb-like transcription factor family protein | ||||
| Swissprot | Q9LVS0 | 0.0 | KUA1_ARATH; Transcription factor KUA1 | ||||
| TrEMBL | R0EUD4 | 0.0 | R0EUD4_9BRAS; Uncharacterized protein | ||||
| STRING | AT5G47390.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3426 | 27 | 59 | Representative plant | OGRP394 | 16 | 99 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G47390.1 |
| Entrez Gene | 834786 |
| iHOP | AT5G47390 |
| wikigenes | AT5G47390 |



