 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016651837.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 336aa MW: 37951.1 Da PI: 5.3161 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.3 | 6.1e-19 | 57 | 110 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
XP_016651837.1 57 KKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 110
56689************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 131.2 | 3.9e-42 | 56 | 148 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrls eqvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+d+ +Lk++yd+lk + ++L++e+e+L +e+k+
XP_016651837.1 56 EKKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDFGVLKANYDSLKLNYDSLQHENEALVKEIKQ 148
69***************************************************************************************9987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.07E-19 | 42 | 114 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.572 | 52 | 112 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.1E-18 | 55 | 116 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 3.9E-16 | 57 | 110 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 4.78E-16 | 57 | 113 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-19 | 59 | 117 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 9.8E-6 | 83 | 92 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 87 | 110 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 9.8E-6 | 92 | 108 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.1E-17 | 112 | 154 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 336 aa Download sequence Send to blast |
MKRLGSSDSL GAMISICPST EEHSPRNNHV YRRDFHSMLD GLDEEGCVEE GGHVSEKKRR 60 LSVEQVKALE KNFEVENKLE PERKVKLAQE LGLQPRQVAV WFQNRRARWK TKQLERDFGV 120 LKANYDSLKL NYDSLQHENE ALVKEIKQLK SKLQEENTES NNLSVKEEQM VAKDQSNYKV 180 VDHEKSKSPP PPPPGSSVPA TESKELNFES FNNTNNGAVG LEAVSLFPDF KDGSSDSDSS 240 AILNEDNSPN LTISSSGMLQ NHQLMKSPAS TSLKFNCSSS SSPSSSSMNC FQFQKTYHPQ 300 FVKIEEHNFF SSEEACSFFS DEQAPTLQWC CPDQWN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 104 | 112 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that may act as growth regulators in response to water deficit. Interacts with the core sequence 5'-CAATTATTA-3' of promoters in response to ABA and in an ABI1-dependent manner. Involved in the negative regulation of the ABA signaling pathway. {ECO:0000269|PubMed:10527431, ECO:0000269|PubMed:12065416}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
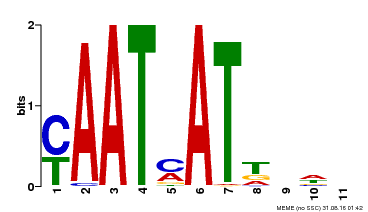
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_016651837.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By water deficit, by abscisic acid (ABA) and by salt stress. Self expression regulation. {ECO:0000269|PubMed:10527431, ECO:0000269|PubMed:12065416, ECO:0000269|PubMed:16055682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016651837.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-6-like isoform X2 | ||||
| Swissprot | P46668 | 6e-75 | ATHB6_ARATH; Homeobox-leucine zipper protein ATHB-6 | ||||
| TrEMBL | M5WIS1 | 0.0 | M5WIS1_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008235150.1 | 0.0 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 5e-66 | homeobox protein 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



