 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf02778g00033.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1261aa MW: 141559 Da PI: 8.9635 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.7 | 0.00038 | 1145 | 1170 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sFs+k +L H ++ +
Peinf101Scf02778g00033.1 1145 YQCDleGCSMSFSSKQELLLHKKNvC 1170
99*******************99866 PP
| |||||||
| 2 | zf-C2H2 | 10.9 | 0.0014 | 1169 | 1192 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp C+k F ++ +L++H r+H
Peinf101Scf02778g00033.1 1169 VCPveGCKKKFFSHKYLVQHRRVH 1192
69999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 12.2 | 0.00056 | 1228 | 1254 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C+ Cg++F+ s++ rH r+ H
Peinf101Scf02778g00033.1 1228 YVCTetGCGQTFRFVSDFSRHKRKtgH 1254
99********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.1E-14 | 17 | 58 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.384 | 18 | 59 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 9.0E-15 | 19 | 52 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 31.429 | 186 | 359 | IPR003347 | JmjC domain |
| SMART | SM00558 | 2.0E-49 | 186 | 359 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 6.59E-26 | 200 | 375 | No hit | No description |
| Pfam | PF02373 | 4.8E-36 | 219 | 342 | IPR003347 | JmjC domain |
| SMART | SM00355 | 7.4 | 1145 | 1167 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.9E-5 | 1166 | 1196 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.572 | 1168 | 1197 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.72E-5 | 1168 | 1205 | No hit | No description |
| SMART | SM00355 | 0.012 | 1168 | 1192 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1170 | 1192 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-9 | 1197 | 1222 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.614 | 1198 | 1227 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1198 | 1222 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1200 | 1222 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.45E-10 | 1208 | 1250 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.8E-10 | 1223 | 1250 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.053 | 1228 | 1259 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.19 | 1228 | 1254 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1230 | 1254 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1261 aa Download sequence Send to blast |
MAEVNIEVFP WLKTLPVAPE YHPTLEEFQD PIAYIFKIEK HASQFGICKI VPPVPPLAKK 60 TILSNLNRSL SARPGSTGPT FTTRQQQIGF CPRKHRPVKK PVWQSGETYT VAEFQGKAKN 120 FEKNYLKKNC FLKKTAALSP LEIETLYWKA TVDKPFSVEY ANDMPGSAFP PKKIGGNGGG 180 GGEGVSLADT EWNMRGVSRA KGSLLKFMKE EIPGVTSPMV YLAMMFSWFA WHVEDHDLHS 240 LNYMHMGAGK TWYGVPRDAA VAFEEFIVKQ YKALFYNVRE YSTITFATLG EKTTVMSPEV 300 LLSAGIPCCR LVQNAGEFVV TFPRAYHSGF SHGFNCGEAA NIATPEWLRF AKEAAIRRAS 360 TNCPPMVSHF QLLYDLALSL CSRVPKNIKI EPRSSRLKDK KRSEGDMLVK ELFVEDLNCN 420 NYLLHILGEG SPVVLLPQDY TGTSNSSYLV AGSQLKVNSR SFPSLSSRDH EVKSLKDSAS 480 DGLMLGRKRG MKQLPRGSLE KGKYSSWHAG NRMPDSGRND DAQSSPETKR GNLDTAREMA 540 YKCDTLSEHG LFSCVTCGIL CYTCVAIVQP TEEAAHHLIS SDFRNFNDWT GDVGGVTAIG 600 RDLNAADSDS SSGWLVKRSP GGLIDVPIES SDRIRKLNNE SVGVLSSTKA RKETSSLGLL 660 ALNYANSSDS DEDEVEADIP VEACESIHTD SEDEVSLRVI DPYANHRQKR AVSEGRICQR 720 FDNPSEYSPS GESNTVSDRL RHHPRSHQVA ANCTPFSHKE EMNNNNDVAP FDNMPMQFTS 780 TSDEDSFRIH VFCLQHAVQV EEQLRQVGGV RISLLCHPDY PKLEAQAKKV AEELGSDHFW 840 REISFREATK EDEEMIQSAL EIEEAIHGNG DWTVKLDMNL FYSANLSRSP LYSKQMPYNF 900 IIYSAFGRSS PDNTPEKSEY TGRGSGKQKR GVVAGKWCGK VWMSSQVHPL LVERDTDEEQ 960 EQNKSIPVQI KPDVKSERPS ERAGTRTVAT TCKTGRKRKS AAEDRNTSNH KLLIADDLDD 1020 SLLSYIPQQH RKTNLRSKRI KYETPEPQQD VDKKKRFSTP SDDELDGGPS TRLRKRIPKP 1080 SKESAAKLVK AKSAIKQHEG MKAEKGSKVK IPSTNRNTKK DPVMKAPRSN TGNTIKKMKD 1140 KEGEYQCDLE GCSMSFSSKQ ELLLHKKNVC PVEGCKKKFF SHKYLVQHRR VHMDDRPLKC 1200 PWKGCKMTFK WAWARTEHIR VHTGERPYVC TETGCGQTFR FVSDFSRHKR KTGHVSKKGR 1260 G |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 5e-71 | 11 | 392 | 8 | 366 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 5e-71 | 11 | 392 | 8 | 366 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 485 | 492 | GRKRGMKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
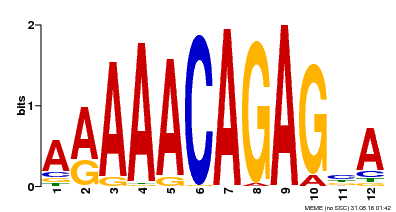
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006351452.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A2G3CCS7 | 0.0 | A0A2G3CCS7_CAPCH; Lysine-specific demethylase REF6 | ||||
| STRING | PGSC0003DMT400039224 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4988 | 23 | 31 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



