 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peinf101Scf01349g03025.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia; Petunia integrifolia
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 691aa MW: 78127.4 Da PI: 8.0031 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 20.9 | 9.3e-07 | 63 | 84 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg sF+ + +Lk+H+++H
Peinf101Scf01349g03025.1 63 TCEECGVSFKKPAHLKQHMQSH 84
6*******************99 PP
| |||||||
| 2 | zf-C2H2 | 16.8 | 1.9e-05 | 90 | 114 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ dC s++rk++L+rH+ H
Peinf101Scf01349g03025.1 90 FVCHvdDCQSSYRRKDHLTRHLLQH 114
89*******************9887 PP
| |||||||
| 3 | zf-C2H2 | 10.8 | 0.0015 | 119 | 144 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+kCp C+ +Fs + ++ +H ++ H
Peinf101Scf01349g03025.1 119 FKCPvdGCKSTFSFQGSMSQHVKKiH 144
89*********************999 PP
| |||||||
| 4 | zf-C2H2 | 20.5 | 1.3e-06 | 176 | 200 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y+Cp Cgk+F+ s L++H +H
Peinf101Scf01349g03025.1 176 YVCPeaGCGKVFKYASKLRKHEDSH 200
9********************9888 PP
| |||||||
| 5 | zf-C2H2 | 19.4 | 2.8e-06 | 296 | 320 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ dC +Fs++snL++Hi+ H
Peinf101Scf01349g03025.1 296 KCEfqDCQHTFSTRSNLVQHIKAaH 320
79999****************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 33 | 15 | 37 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 4.4E-10 | 60 | 114 | No hit | No description |
| PROSITE profile | PS50157 | 13.547 | 62 | 89 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0033 | 62 | 84 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.9E-8 | 63 | 90 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 64 | 84 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.411 | 90 | 119 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.035 | 90 | 114 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-9 | 91 | 116 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 92 | 114 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.8E-5 | 100 | 145 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.4E-4 | 117 | 145 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 8.954 | 119 | 144 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0048 | 119 | 144 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 121 | 144 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-8 | 173 | 202 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.97 | 176 | 205 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 4.3E-4 | 176 | 200 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 178 | 200 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 9.3 | 237 | 262 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 239 | 262 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 13 | 265 | 286 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 8.5E-6 | 276 | 317 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 6.14E-9 | 277 | 333 | No hit | No description |
| PROSITE profile | PS50157 | 13.214 | 295 | 325 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0037 | 295 | 320 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 297 | 320 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 6.6E-5 | 319 | 348 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 11 | 326 | 352 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 328 | 352 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS51375 | 5.042 | 404 | 441 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 5.437 | 442 | 476 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.32 | 477 | 507 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 0.0011 | 483 | 508 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF13041 | 4.1E-12 | 507 | 548 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 13.417 | 508 | 542 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 8.8E-9 | 510 | 543 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.966 | 545 | 579 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 0.001 | 547 | 573 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 5.831 | 611 | 641 | IPR002885 | Pentatricopeptide repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 691 aa Download sequence Send to blast |
MGEEEEVKKR DIRRYYCEYC GLCRSKKSLI SSHILSHHQD EIERRKDEGN EDANVKEGPK 60 MNTCEECGVS FKKPAHLKQH MQSHSIERPF VCHVDDCQSS YRRKDHLTRH LLQHQGKLFK 120 CPVDGCKSTF SFQGSMSQHV KKIHDPSVSP KXNMSQHVKK IHDPSVSPKV DPPKQYVCPE 180 AGCGKVFKYA SKLRKHEDSH GKSNLKILIF LLICVLKTSN NIKLPCIYAV KLETVEALCL 240 EPGCMKHFTN EKCLKEHINT CHQHIVCEIC GTKQLKKNIK RHLRTHEEGP TSERVKCEFQ 300 DCQHTFSTRS NLVQHIKAAH LGDKPFSCGI PGCGMKFAFK HVRDRHEKSG CHVYTPGDFV 360 EADEQFRSRP RGGRKRKLPV FEDIMRKRIT PPSGTDRVFN QGSEYLSWLL SVESDDELVC 420 SNTAREAVEL FVEMRNEGFE PDERTLVSVL GACGDLGDVN LGKWVEEYVS GKNMELNSFM 480 GSALINMYGK CGDLVSARRI FDGMRKKDVI TWNAMISGYA QNGLSDETVS MFNAMKESGV 540 DPNKITISWN AMISAFAFHG RAQEALSLFE RMTFESSDAC PDDVTFVGVL SACGWLIKAG 600 IMSSSFGLVP KVEHFSCIVY LLSRAGCVYE SWEFTEKMPQ KPDGILLDAL VGAYRKLKNI 660 DVGEQVMQLL LVMEPSNSGN YIISSKIYAD L |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5iww_D | 3e-29 | 416 | 666 | 25 | 330 | PLS9-PPR |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 368 | 376 | RPRGGRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
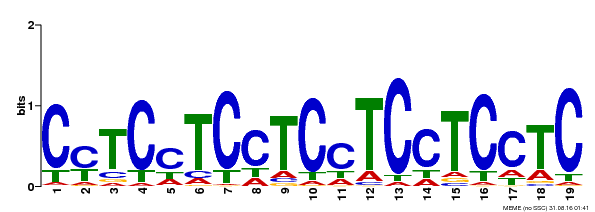
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016453639.1 | 0.0 | PREDICTED: transcription factor IIIA-like isoform X2 | ||||
| Swissprot | Q84MZ4 | 1e-135 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A1S3YMZ9 | 0.0 | A0A1S3YMZ9_TOBAC; transcription factor IIIA-like isoform X2 | ||||
| STRING | XP_009592258.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3644 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-129 | transcription factor IIIA | ||||



