 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01001451G0080 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 645aa MW: 70108.6 Da PI: 5.9504 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 96.3 | 2.8e-30 | 88 | 172 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+++e+laLi++r+em+ ++r++ lk+p Weevs+k++e g++rs+k+Ck k+en+ k+yk++keg+ +r++++s +++f+qlea
PH01001451G0080 88 RWPREETLALIRIRSEMDAAFRNADLKAPVWEEVSRKLAELGYHRSAKKCKGKFENVDKYYKRTKEGRAGRQDGKS--YRFFSQLEA 172
8*********************************************************************866665..*******85 PP
| |||||||
| 2 | trihelix | 109.1 | 2.8e-34 | 393 | 478 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k+ev+aLi++r e++e++++ k+plWe++s+ mr+ g++rs+k+Ckekwen+nk++kk+ke++k+r +e+s+tcpyf+ql+a
PH01001451G0080 393 RWPKEEVQALIQLRMEKDEQYQDVGPKGPLWEDISAGMRRIGYNRSAKRCKEKWENINKYFKKVKESNKRR-PEDSKTCPYFHQLDA 478
8********************************************************************97.99***********85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 6.829 | 81 | 145 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.02 | 85 | 147 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 2.35E-26 | 87 | 152 | No hit | No description |
| Pfam | PF13837 | 1.4E-21 | 87 | 173 | No hit | No description |
| SMART | SM00717 | 1.9E-5 | 390 | 452 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 7.724 | 392 | 450 | IPR017877 | Myb-like domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-4 | 392 | 449 | IPR009057 | Homeodomain-like |
| CDD | cd12203 | 8.01E-28 | 392 | 457 | No hit | No description |
| Pfam | PF13837 | 3.5E-22 | 392 | 479 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 645 aa Download sequence Send to blast |
MQHHQGSAGS PYRVPTTDMV PFSLPPASGA VSVSSTPPPM QPQPAGTSFE ELPAGGSAAA 60 AVNFHDDYML VDVGCGRSVS KAGGSGNRWP REETLALIRI RSEMDAAFRN ADLKAPVWEE 120 VSRKLAELGY HRSAKKCKGK FENVDKYYKR TKEGRAGRQD GKSYRFFSQL EALHAAAPQQ 180 HQGTAMATAA YTATVQDPQP LAMARMAPAV LAHPGASLPD LGFSSNSESE SDDESGDEEE 240 EGPGKLRSGD GGSNKRMMAF FQGMMKQVTE KQDAMHRLFL ETLDKWEAER TARDETWRRK 300 EVARINRERE ELAKERTAAA SRDAALFAFL KRVGGQSVQL PPSGTVAVPT PMPAHAPPPP 360 HHDAAATSLQ LVPVQPKVEE GWAGGESCGA TSRWPKEEVQ ALIQLRMEKD EQYQDVGPKG 420 PLWEDISAGM RRIGYNRSAK RCKEKWENIN KYFKKVKESN KRRPEDSKTC PYFHQLDAIY 480 RKKHFAGSGG GSSAVSGASM AAVTASEKEN PQCEFELKSS NDFDKRSSGG GQNVQAPPGN 540 GETAPATTVR DSATNKEAEE TLTMEPNVQS QQQEFTAPDE TDSDDMEGNY TDEGDDDGNE 600 DNRMQHKIEF QKPKESGSSS APGPATTAAV TSSTPTSSTF LAVQ* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
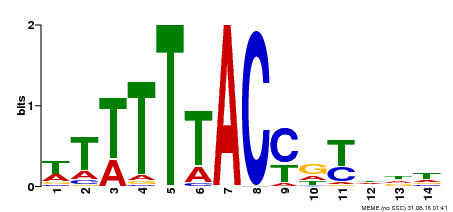
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01001451G0080 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK375494 | 1e-119 | AK375494.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3095N08. | |||
| GenBank | AK376129 | 1e-119 | AK376129.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3115F22. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004953287.1 | 0.0 | trihelix transcription factor GTL1 isoform X1 | ||||
| Refseq | XP_012699251.1 | 0.0 | trihelix transcription factor GTL1 isoform X1 | ||||
| Swissprot | Q39117 | 1e-113 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | A0A287UDK8 | 0.0 | A0A287UDK8_HORVV; Uncharacterized protein | ||||
| TrEMBL | A0A287UDK9 | 0.0 | A0A287UDK9_HORVV; Uncharacterized protein | ||||
| STRING | Si016578m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP606 | 38 | 175 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76880.1 | 1e-103 | Trihelix family protein | ||||



