 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000667G0350 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 481aa MW: 51581.7 Da PI: 6.267 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 88.7 | 5.3e-28 | 182 | 236 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k +++WtpeLH+rFv+aveqL G++kA+P++ilelm+v++Lt+++++SHLQkYR++
PH01000667G0350 182 KVKVDWTPELHRRFVQAVEQL-GIDKAVPSRILELMGVECLTRHNIASHLQKYRSH 236
5689*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.86 | 179 | 238 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.3E-27 | 180 | 239 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.05E-18 | 180 | 239 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.8E-27 | 182 | 236 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.1E-7 | 185 | 234 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 481 aa Download sequence Send to blast |
MLEVSTLRSP KADQRAVVGD HHVGFPASSA TDDDDFIVDD NLLDYIDFSC DVPFFHADGD 60 ILPDLEVDPT ELLAEFSSPD KPIPTVSPAD PAADGCGARG DAGAEAKTTE VKEGKKELIE 120 EKDEKGFLEE KDVKSNGDEV LSAVTTEDSA AVGSDTKSSA SAEGHSKRKP SSSAKNSQGK 180 RKVKVDWTPE LHRRFVQAVE QLGIDKAVPS RILELMGVEC LTRHNIASHL QKYRSHRKHL 240 MAREAEAASW TQKRPIYVAA AAGTGGQVAG GPREDVAGPW VVPTIGFPPP GMPHAVAAMV 300 QPPPPPCCRP LHVWGHPTAV EVAPPPMLPV WPRHLAPPQP LPAWAHQPPV DPAYWHQQYN 360 AARKWGPQSV TQGTPCIPPP MAPAAMLQRF PLPPMPGIFP HPMYRPLPPP QSNKLAGLQL 420 QLDAHPSKES IDAAIGDVLV KPWLPLPLGL KPPSLDSVMS ELQKQGVPKV PPAATTSGGA 480 G |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5lxu_A | 1e-15 | 187 | 238 | 6 | 57 | Transcription factor LUX |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that promotes chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport (By similarity). {ECO:0000250}. | |||||
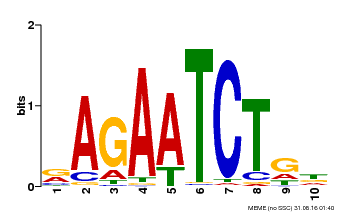
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000667G0350 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. {ECO:0000269|PubMed:11340194}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FP099309 | 0.0 | FP099309.1 Phyllostachys edulis cDNA clone: bphyem212o08, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020187877.1 | 0.0 | probable transcription factor GLK2 | ||||
| Swissprot | Q5NAN5 | 1e-156 | GLK2_ORYSJ; Probable transcription factor GLK2 | ||||
| TrEMBL | A0A0A9LLL4 | 0.0 | A0A0A9LLL4_ARUDO; Uncharacterized protein | ||||
| STRING | ORUFI01G09620.1 | 1e-156 | (Oryza rufipogon) | ||||
| STRING | Si001336m | 1e-156 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2415 | 38 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 5e-74 | GBF's pro-rich region-interacting factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



