 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PH01000201G0710 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Bambusoideae; Arundinarodae; Arundinarieae; Arundinariinae; Phyllostachys
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1264aa MW: 141064 Da PI: 6.3979 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 176.7 | 3e-55 | 240 | 356 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
ke ++rwl++ ei++iL+n+++++++ e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+e+n++fq
PH01000201G0710 240 KEaHQRWLRSAEICEILKNYRNFRIAPEPPNRPSSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKSGSIDVLHCYYAHGEDNENFQ 333
5559****************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+Lee++ +ivlvhyle+k
PH01000201G0710 334 RRTYWMLEEDFMHIVLVHYLETK 356
********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 80.852 | 235 | 361 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.4E-75 | 238 | 356 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 7.0E-49 | 241 | 354 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 1.7E-6 | 683 | 770 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.3E-6 | 684 | 762 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 3.2E-17 | 684 | 770 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 5.6E-15 | 876 | 981 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 9.2E-16 | 879 | 980 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.53E-11 | 879 | 977 | No hit | No description |
| PROSITE profile | PS50297 | 15.671 | 885 | 979 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.736 | 918 | 950 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 6.37E-8 | 1078 | 1131 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.4 | 1080 | 1102 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.096 | 1081 | 1110 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.034 | 1082 | 1101 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 1103 | 1125 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.56 | 1104 | 1128 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.5E-4 | 1106 | 1125 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1264 aa Download sequence Send to blast |
MGPFQVVYGF NPRTPIDLLP IPSSKKVNLD AKKRVEFVLK LHETTKENIE RMNARYKIAA 60 SHGRKSLTFE LGDLIWLHLR KDRFPELRKS KLMPRADGPF KVLAKINDNA YKLDLPAEFR 120 VSLTFNIAGI DPYDQIGTTR VHHTKTLGLA AVASYFKLMS LPLDEVVHCN RSSCGGPLAR 180 SDWSSSDLAR PRPEEVGRPM GTVGWRKSPV VAEEQAFLSL EDDWALGIGV LLGDIEQILK 240 EAHQRWLRSA EICEILKNYR NFRIAPEPPN RPSSGSLFLF DRKVLRYFRK DGHNWRKKKD 300 GKTVKEAHER LKSGSIDVLH CYYAHGEDNE NFQRRTYWML EEDFMHIVLV HYLETKGGKS 360 SRARGNNDML QAAVVDSPLS QLPSQTLEGE SSLSGQASEY EDAESVSLIH IPIMHSVSKA 420 KFGAADTYSG GAGYYSFTQM QQHQNGTGPV IDASIFSPYA PASSIGNYQG LQHAIAHNTS 480 FYSSSQDNSP VVLNDSSVGL AMNGRESQTD LPSWNEVMKP DKGSVQMPRL QLPVHPEEGT 540 FTEGPGVEYL TFDEVYSDGL SLNDFSADGE SPWQFSSATG DLSSTENNFP QNDGSLEAAI 600 GYPFLKSQSS NLSDILKDSF KKTDSFTRWM SKELPEVEDS QIQSSSGAYW NTEEADSIIE 660 ASSREPLDQF TVAPMLSQDQ LFSIVDFSPS WTYTGSKIKV LVTGKFLNTN QVTERCKWSC 720 MFGEVEVPAE ISADGTLHCY SPPHKPGRVP FYITCSNRLA CSEVREFEFR PSPSQYMDAP 780 SPHGATIKVY FQIRLDKLLS LVPDEYQATI SNPSLEMIDL SKKISSLMTK NDEWSKLLKL 840 ADDNELLTDD QQDQFAENLI KEKLHVWLLH KVGDGGKGPS VLDDEGQGVL HLAAALGYDW 900 AIRPTVTAGV NINFRDVHGW TALHWAAFCG RERTVVALIA LGAAPGALTD PTPDFPSVST 960 PADLASANGH KGISGFLAES YLTSHLQVLN LKEASMAEIS GLPGIGDVTE RSASQPAIGD 1020 SLDAVRNAAQ AAARIYQVFR VQSFQRKQAV QYEDEKSGIS DERALSLLSV KPSKPGQLDP 1080 LHAAASRIQN KFRGWKGRKE FLLIRQRIVK IQAHVRGHQV RKHYRKIVWS VGIVEKVILR 1140 WRRRGAGLRG FRSAEAAIES SSCGTSSNLV ADKPAADDYD FLQEGRKQTE ERLQKALARV 1200 KSMAQYPEAR DQYQRILTVV SKMQESQAMQ EKMLEDSTEM DEGYFMSEFK ELWNDDTPLP 1260 DYF* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
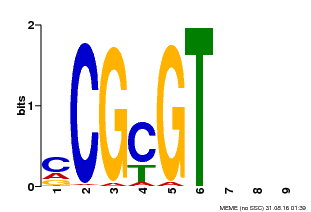
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PH01000201G0710 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK101479 | 0.0 | AK101479.1 Oryza sativa Japonica Group cDNA clone:J033041A20, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010234691.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A3L6TUJ2 | 0.0 | A0A3L6TUJ2_PANMI; Calmodulin-binding transcription activator 3-like isoform X2 | ||||
| STRING | Pavir.Ia02749.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



