 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.C04225.3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1137aa MW: 127639 Da PI: 7.7748 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.7 | 0.00019 | 1017 | 1040 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
++C C++sFs++ +L H r
Pahal.C04225.3 1017 FSCDieGCDMSFSTQQDLALHKRD 1040
899999***************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 9.7 | 1017 | 1039 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.887 | 1040 | 1069 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 1040 | 1064 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 9.92E-5 | 1041 | 1073 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 1042 | 1064 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-9 | 1067 | 1094 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.344 | 1070 | 1099 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.002 | 1070 | 1094 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1072 | 1094 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.08E-9 | 1080 | 1123 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.2E-9 | 1095 | 1123 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.845 | 1100 | 1131 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.1 | 1100 | 1126 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1102 | 1126 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1137 aa Download sequence Send to blast |
MRRQLSCLLK CFFLLEFLAA DWYKTLVNLL SRSLELIIPV LAMVAKEAAV RRASTNCGPL 60 VSHYQLLYEL ALSLRPRELK NSHDVPRSSR LRDKKKNESE IMIKETFVGS VIENNSFLSI 120 LLDKSSCVII PEIEFPLPSF PTMMVPEVTV KQALIAGPCS IRQKKAEDML ASATTSFAFN 180 GRKLYETKFG TVNSSAFLLN PEIQSGVIEK GRSHQGGGLL DQGRLPCVQC GILSYACVAI 240 IQPKEAAVQY VISQECMSSS AKHGEIMKSD DTPNWISIVP PQGHSSETDD NTIHNVNSAH 300 VSDRCRQLYT SSTHGCTSAL GLLASAYDSS DSDEEAEMPN EIANISANND AENGVTNIQS 360 SGRSIQHQNT NLHLSEEECD PRATPSPMKP VDDRSIAMTQ ASIGTDMTRL AELGESLTAY 420 EQWSPYVDLD DDPTASGAKT SLNTSFSRAK GAMEPDALTL LKYSKDSCRM HVFCLEHALE 480 TWTQLQQIGG ANIMLLCHPE YPAAELAAKV IAEELGMKHA WKDITFKQAT DEDIGRIRLA 540 LQDEDAEPTS SDWAVKMGIN IYYSAKQSKS PLYSKQVPYN SIIYKAFAQE NPDRDEERQR 600 SRTTKKKVAG SWCGKVWMSN QVHPLLACER EEEDLDMVCS KAMVPVSSYD RIQEEPSTRS 660 TSLISRSLSK RISRRKEVDS VEKSRAKKKR YTTSDLATFD QPRNCDDHDK HEDGGESESE 720 DAQNTQQHQQ YESQKMNKKS SSKRRKDDKR KNYFYERRNY NDDIDYKLSI DWDNTPPQGL 780 DVVEVKSGAQ LQGSKKKSSK CKANDDLSNV EKKLQKMGKK VSTKKHKNDK TNQQFQGNHN 840 EDNVDLLPED NGDEASQESW DEVPTQKTDD VKVKSRGKMH SGKKKASKCQ TSDGLDNVDL 900 LHEDNGYEAT QESWDEVPKQ KTDDVRVNSR GKTHIGKKKA SKCQISDGLD NGDNEAKFSC 960 DTAVCNRDKA TIDDWEEIPK EKADDVKVKS NMQSGKKKAS KRPASDGLRN GDKGAKFSCD 1020 IEGCDMSFST QQDLALHKRD ICPVKGCKKK FFCHKYLLQH RKVHLDERPL MCSFTGCKKT 1080 FKWPWARTEH MRVHTGVRPY ACTEPGCTQT FRFVSDFSRH KRKTGHSSDK KKKNST* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 3e-54 | 1015 | 1133 | 21 | 139 | Lysine-specific demethylase REF6 |
| 6a58_A | 3e-54 | 1015 | 1133 | 21 | 139 | Lysine-specific demethylase REF6 |
| 6a59_A | 3e-54 | 1015 | 1133 | 21 | 139 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 673 | 689 | RRKEVDSVEKSRAKKKR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
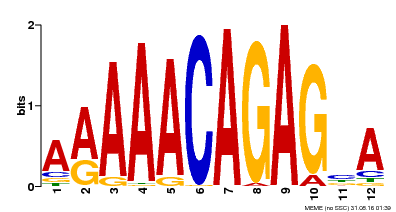
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025807104.1 | 0.0 | lysine-specific demethylase JMJ705-like | ||||
| TrEMBL | A0A2S3HD45 | 0.0 | A0A2S3HD45_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Cb01041.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 4e-53 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.C04225.3 |



