 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.C04225.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1449aa MW: 161426 Da PI: 7.3184 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.3 | 0.00025 | 1329 | 1352 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
++C C++sFs++ +L H r
Pahal.C04225.1 1329 FSCDieGCDMSFSTQQDLALHKRD 1352
899999***************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.7E-16 | 17 | 58 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.451 | 18 | 59 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 8.3E-14 | 19 | 52 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 35.512 | 196 | 365 | IPR003347 | JmjC domain |
| SMART | SM00558 | 3.2E-46 | 196 | 365 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 2.75E-24 | 210 | 361 | No hit | No description |
| Pfam | PF02373 | 4.0E-36 | 229 | 348 | IPR003347 | JmjC domain |
| SMART | SM00355 | 9.7 | 1329 | 1351 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.887 | 1352 | 1381 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 1352 | 1376 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1354 | 1376 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-9 | 1379 | 1406 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.344 | 1382 | 1411 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.002 | 1382 | 1406 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1384 | 1406 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.07E-9 | 1392 | 1435 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 4.2E-9 | 1407 | 1435 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.1 | 1412 | 1438 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.845 | 1412 | 1443 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1414 | 1438 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1449 aa Download sequence Send to blast |
MSPPAVETPE WLRNLPVAPE YRPTAAEFAD PIAYILKIEA EASRYGICKV VPPLAAPPRE 60 ATVERLRASF AANAAAGSGI DGATPAPTFP TRLQQVGFST KNRRPASRRV WESGERYTLE 120 AFRAKARDIE LPRHAVPPKH ATQLQLEALF WGACAARPFN VEYGNDMPGS GFAAPEELDL 180 DLEGGGGGNA ALAARDVGET EWNMRLAPRA RGSLLRAMGR DVAGVTTPML YVAMLYSWFA 240 WHVEDHELHS LNYLHFGKPK TWYGVPRDAM LAFEDAVRVH GYADDLNAIM AFQTLNEKTT 300 VLSPEVLLSA GVPCCRLVQN PGEFIITFPG AYHSGFSHGF NCGEATNIAT PRWLQVAKEA 360 AVRRASTNCG PLVSHYQLLY ELALSLRPRE LKNSHDVPRS SRLRDKKKNE SEIMIKETFV 420 GSVIENNSFL SILLDKSSCV IIPEIEFPLP SFPTMMVPEV TVKQALIAGP CSIRQKKAED 480 MLASATTSFA FNGRKLYETK FGTVNSSAFL LNPEIQSGVI EKGRSHQGGG LLDQGRLPCV 540 QCGILSYACV AIIQPKEAAV QYVISQECMS SSAKHGEIMK SDDTPNWISI VPPQGHSSET 600 DDNTIHNVNS AHVSDRCRQL YTSSTHGCTS ALGLLASAYD SSDSDEEAEM PNEIANISAN 660 NDAENGVTNI QSSGRSIQHQ NTNLHLSEEE CDPRATPSPM KPVDDRSIAM TQASIGTDMT 720 RLAELGESLT AYEQWSPYVD LDDDPTASGA KTSLNTSFSR AKGAMEPDAL TLLKYSKDSC 780 RMHVFCLEHA LETWTQLQQI GGANIMLLCH PEYPAAELAA KVIAEELGMK HAWKDITFKQ 840 ATDEDIGRIR LALQDEDAEP TSSDWAVKMG INIYYSAKQS KSPLYSKQVP YNSIIYKAFA 900 QENPDRDEER QRSRTTKKKV AGSWCGKVWM SNQVHPLLAC EREEEDLDMV CSKAMVPVSS 960 YDRIQEEPST RSTSLISRSL SKRISRRKEV DSVEKSRAKK KRYTTSDLAT FDQPRNCDDH 1020 DKHEDGGESE SEDAQNTQQH QQYESQKMNK KSSSKRRKDD KRKNYFYERR NYNDDIDYKL 1080 SIDWDNTPPQ GLDVVEVKSG AQLQGSKKKS SKCKANDDLS NVEKKLQKMG KKVSTKKHKN 1140 DKTNQQFQGN HNEDNVDLLP EDNGDEASQE SWDEVPTQKT DDVKVKSRGK MHSGKKKASK 1200 CQTSDGLDNV DLLHEDNGYE ATQESWDEVP KQKTDDVRVN SRGKTHIGKK KASKCQISDG 1260 LDNGDNEAKF SCDTAVCNRD KATIDDWEEI PKEKADDVKV KSNMQSGKKK ASKRPASDGL 1320 RNGDKGAKFS CDIEGCDMSF STQQDLALHK RDICPVKGCK KKFFCHKYLL QHRKVHLDER 1380 PLMCSFTGCK KTFKWPWART EHMRVHTGVR PYACTEPGCT QTFRFVSDFS RHKRKTGHSS 1440 DKKKKNST* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 3e-68 | 10 | 386 | 7 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 3e-68 | 10 | 386 | 7 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 985 | 1001 | RRKEVDSVEKSRAKKKR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
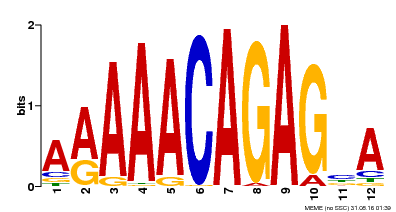
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025807104.1 | 0.0 | lysine-specific demethylase JMJ705-like | ||||
| TrEMBL | A0A2S3HD45 | 0.0 | A0A2S3HD45_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Cb01041.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6718 | 30 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-158 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.C04225.1 |



