 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.C01957.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 729aa MW: 79471 Da PI: 7.4986 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 77.9 | 1.1e-24 | 159 | 260 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f+k+lt sd++++g +++p++ ae++ + +++ s++l+ +d++g +W++++iyr++++r++lt+GW+ Fv+ ++L +gD v+F + ++
Pahal.C01957.2 159 FCKTLTASDTSTHGGFSVPRRAAEDCfppldyS-QQRPSQELVAKDLHGTEWRFRHIYRGQPRRHLLTTGWSAFVNKKKLVSGDAVLFL--RGDD 250
99****************************963.3445679************************************************..3489 PP
E..EEEEE-S CS
B3 90 felvvkvfrk 99
+el+++v+r+
Pahal.C01957.2 251 GELRLGVRRA 260
999***9996 PP
| |||||||
| 2 | Auxin_resp | 109.8 | 2.9e-36 | 286 | 365 | 2 | 82 |
Auxin_resp 2 ahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsL 82
aha++tksvF+++YnPr s+seF+++++k++k++++++s+G Rfkm++e++d+ err++G+++g+ d+dp W++SkW++L
Pahal.C01957.2 286 AHAVATKSVFHIYYNPRLSQSEFIIPYSKFMKSFSQPFSAGLRFKMRYESDDATERRYTGIIAGIGDADP-MWRGSKWKCL 365
89********************************************************************.9********9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 1.7E-43 | 150 | 274 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 7.9E-41 | 152 | 262 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 5.56E-23 | 157 | 259 | No hit | No description |
| PROSITE profile | PS50863 | 13.493 | 159 | 261 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 5.3E-23 | 159 | 261 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.3E-22 | 159 | 260 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.1E-30 | 286 | 365 | IPR010525 | Auxin response factor |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009725 | Biological Process | response to hormone | ||||
| GO:0010050 | Biological Process | vegetative phase change | ||||
| GO:0010158 | Biological Process | abaxial cell fate specification | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 729 aa Download sequence Send to blast |
MTGIDLNDTV EEDEAEAEAG NSCSQQSRSS SAATGTPPPP GPGAAVCLEL WHACAGPVAP 60 LPRKGSVVVY LPQGHLEHLG DAAAAAGGGT VPPHVFCRVV DLTLHADAST DEVYAQLALV 120 AENEEVARRL RGGSEDGSGG DGEDGDTVRQ RFSRMPHMFC KTLTASDTST HGGFSVPRRA 180 AEDCFPPLDY SQQRPSQELV AKDLHGTEWR FRHIYRGQPR RHLLTTGWSA FVNKKKLVSG 240 DAVLFLRGDD GELRLGVRRA AQLKNGSAFP ALYNQCSNLG SLANIAHAVA TKSVFHIYYN 300 PRLSQSEFII PYSKFMKSFS QPFSAGLRFK MRYESDDATE RRYTGIIAGI GDADPMWRGS 360 KWKCLMVRWD DDVDFRRPNR ISPWEIELTS SVSGSHLSAP NAKRLKPCLP HVNPDYLVPN 420 GSGRPDFAES AQFHKVLQGQ ELLGYRTRDN DAIATSQPCE ARNMHYIDER SCSNDASNNI 480 PGVPRHGART PLGNPGFPYH CSGFGESQRF QKVLQGQEVF RPYRGTLADA CLRNNAFHQQ 540 DGSHAPSVAN KWHTQLHGCA FRGSPAPMNP SQSSSPPSVL IFQQGNSKLS QFEFGHGPLD 600 KNVDGRPAML GHAGGIGGTE QTLMLQPHHV SEEVGNRHAI VEKFHSVGKD GTDNRDVNTN 660 SCKIFGISLA EKVPSSKQKE CGDANYPSPF LSLKQQVPKS LGNSCATVHE QRPVVGRVID 720 LSTMDMMI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-112 | 45 | 395 | 18 | 361 | Auxin response factor 1 |
| 4ldw_A | 1e-112 | 45 | 395 | 18 | 361 | Auxin response factor 1 |
| 4ldw_B | 1e-112 | 45 | 395 | 18 | 361 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00021 | PBM | Transfer from AT2G33860 | Download |
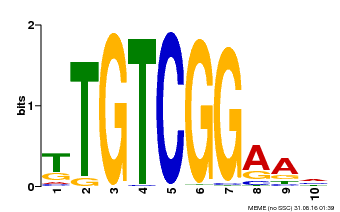
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025807170.1 | 0.0 | auxin response factor 15-like | ||||
| Swissprot | Q8S985 | 0.0 | ARFO_ORYSJ; Auxin response factor 15 | ||||
| TrEMBL | A0A2S3H9P0 | 0.0 | A0A2S3H9P0_9POAL; Auxin response factor | ||||
| STRING | Pavir.J08401.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33860.1 | 1e-165 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.C01957.2 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



