 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.C01148.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 375aa MW: 42670.6 Da PI: 8.7436 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 21.8 | 4.9e-07 | 75 | 97 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ Cg++F+ +s+Lk+H+++H
Pahal.C01148.2 75 HDCKECGMRFKKPSHLKQHMQSH 97
56*******************99 PP
| |||||||
| 2 | zf-C2H2 | 19.2 | 3.3e-06 | 103 | 127 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ C+ C+ s+srk++L+rH+ tH
Pahal.C01148.2 103 FACHvdGCPLSYSRKDHLNRHLLTH 127
78*********************99 PP
| |||||||
| 3 | zf-C2H2 | 14.6 | 9.3e-05 | 132 | 157 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++Cp C++ F+ k n++rH + H
Pahal.C01148.2 132 FVCPieGCDRKFNIKGNMQRHVQEmH 157
89*******************99877 PP
| |||||||
| 4 | zf-C2H2 | 20.8 | 1e-06 | 169 | 193 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp +Cgk+F+ s Lk+H +H
Pahal.C01148.2 169 FICPevNCGKTFKYASKLKKHEESH 193
78*******************9988 PP
| |||||||
| 5 | zf-C2H2 | 16.5 | 2.5e-05 | 260 | 284 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC dC+ sFs ksnL++H + H
Pahal.C01148.2 260 KCRfeDCKCSFSKKSNLEKHVKAvH 284
68888***************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50157 | 8.538 | 29 | 57 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 12 | 29 | 52 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 14.316 | 75 | 102 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0041 | 75 | 97 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.3E-6 | 75 | 94 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 77 | 97 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.32E-11 | 88 | 129 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.6E-14 | 95 | 129 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0053 | 103 | 127 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.328 | 103 | 132 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 105 | 127 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 9.31E-7 | 113 | 163 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 5.1E-8 | 130 | 158 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.533 | 132 | 162 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.9E-4 | 132 | 157 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 134 | 157 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-6 | 165 | 193 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 7.34E-5 | 166 | 195 | No hit | No description |
| SMART | SM00355 | 0.0032 | 169 | 193 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.281 | 169 | 193 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 171 | 193 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 7.3 | 201 | 226 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 203 | 226 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 229 | 250 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 5.39E-6 | 240 | 297 | No hit | No description |
| SMART | SM00355 | 0.041 | 259 | 284 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.03 | 259 | 289 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.6E-4 | 259 | 284 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 261 | 284 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.7E-6 | 285 | 312 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 2.9 | 290 | 316 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.016 | 290 | 319 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 292 | 316 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 375 aa Download sequence Send to blast |
MGDGEETGCG DGATTAARAG APLRDIRRYK CEFCGVVRSK KSLIRAHVLQ NHKDEVDGLE 60 DYQEGGDGAS RKVSHDCKEC GMRFKKPSHL KQHMQSHSLE RPFACHVDGC PLSYSRKDHL 120 NRHLLTHQGK LFVCPIEGCD RKFNIKGNMQ RHVQEMHKDG SPCESKKEFI CPEVNCGKTF 180 KYASKLKKHE ESHVKLDYTE VICCEPGCMK TFTDVECLKA HNQSCHQHVQ CDVCGTKQLK 240 RNFKRHRQMH EGSCITERVK CRFEDCKCSF SKKSNLEKHV KAVHEQRRPF VCQFSGCGKK 300 FSYKHVRDIH EKSSAHVHTE GDFVEADEQR PRSAGGRKRK PVSVETFMRK RVAAPDDAPA 360 HVDGTEYLRW LLSG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 1e-17 | 79 | 223 | 19 | 157 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 1e-17 | 79 | 223 | 19 | 157 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
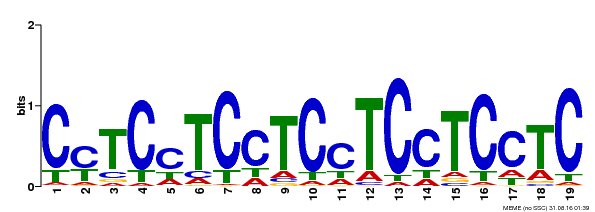
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025805294.1 | 0.0 | transcription factor IIIA-like | ||||
| Refseq | XP_025805295.1 | 0.0 | transcription factor IIIA-like | ||||
| Swissprot | Q84MZ4 | 1e-117 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A2S3H7J2 | 0.0 | A0A2S3H7J2_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.J02126.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-118 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.C01148.2 |



