 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CCG020006.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1081aa MW: 121279 Da PI: 5.2891 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.8 | 6.6e-56 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcy 99
++ ++rwl++ ei++iL n+++++++ e+++ p+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+v+vl+cyYah+e+n++fqrr+y
CCG020006.1 20 VEAQHRWLRPAEICEILSNYQRFRIAPEPAHMPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKSGSVDVLHCYYAHGEDNENFQRRSY 117
6779********************************************************************************************** PP
CG-1 100 wlLeeelekivlvhylevk 118
wlLeeel++ivlvhy+evk
CCG020006.1 118 WLLEEELSHIVLVHYREVK 136
****************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.113 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 8.9E-77 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 5.9E-49 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 1.7E-4 | 503 | 586 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 1.71E-15 | 503 | 588 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 3.3E-4 | 503 | 589 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF48403 | 5.6E-15 | 678 | 794 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 5.5E-16 | 682 | 795 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.90E-11 | 689 | 792 | No hit | No description |
| PROSITE profile | PS50297 | 16.547 | 700 | 804 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.778 | 733 | 765 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.011 | 733 | 762 | IPR002110 | Ankyrin repeat |
| Pfam | PF00023 | 5.4E-4 | 733 | 764 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3300 | 772 | 801 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.09E-7 | 903 | 957 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 2 | 906 | 928 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0045 | 909 | 927 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.206 | 909 | 936 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0039 | 929 | 951 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.597 | 930 | 954 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 932 | 951 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1081 aa Download sequence Send to blast |
MADTRRYPLG NQLDIQQILV EAQHRWLRPA EICEILSNYQ RFRIAPEPAH MPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHER LKSGSVDVLH CYYAHGEDNE NFQRRSYWLL 120 EEELSHIVLV HYREVKQGTR TNFNRIKEHE ECIPYSQETE DTMPSSEMDT SVSSGFHPNG 180 YQVPTRTTDT ASMNSAQASE YEDAESVYNN QASSRFHSFL EVQKPAMEKI DTGSSVHYDH 240 MTFPSDYQGI LSAVPGMDFI SLAQVDKTKE TNGTESACEP QKVTDLPSWD DVLENGACGI 300 ESVPFQTLLS QDDTEGIIPK QDGILEKLLT NSFDKREDIG SHILDQEAWQ SMDGVSSHLL 360 KWSVDQKLLL DSGYDLTARF PDEQLDSGNL INTLEPLCTQ ENDILIQNDQ GMTLEGKPMY 420 SSLVKHHILD GSGTEGLKKL DSFTRWMSKE LGDVEPQVQS SSGSYWITVE SENGVDDSSN 480 PSQGNLDAYL LSPSLSQDQL FSIIDFSPNW AYAGTEIKVL IMGRFLKGRE AAENFQWSIM 540 FGEVEVPAEV IADGVLRCNT PSHKAGRIPF YVTCSNRVAC SEVREFEYLS HTQDITYNYR 600 DSVTEDLNMR FGKLLSLSSV SPSKYDSSSV DEILSSKINS LLNEDNETWD QMFKLTSEEG 660 FSSEKVKERL VQKLLKEQLH VWLLQKASEG GKGPSILDEG GQGVLHFAAA LGYDWALEPT 720 IVAGVSVNFR DVNGWTALHW AASYGRERTV ASLIHLGAAP GALTDPTPTY PTSRTPADLA 780 SANGHKGISG FLAESALSAH LSSLNLEKQD GKAAECSGTQ ASQTVSGCNA TPVNDADLPS 840 GLPLKDSLAA VCNATQAAAR IHQVFRVQSF QKKQLKEYGD DKLGMSHERA LSLIAVKSQK 900 AGQYDEPVHA AIRIQNKFRG WKGRKEFLII RQRIVKIQAH VRGHQVRKNY RKIIWSVGIL 960 DKIILRWRRK GSGLRGFKSE ALTDGSSMQV VQSKDDDDDF LKEGRRQTEE RSQIALARVK 1020 SMHQHPEARE QYCRLRNVVA EIQEAKAMWE WANNSEVMAE FDELVDLGTS MDDDSFMPSN 1080 S |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
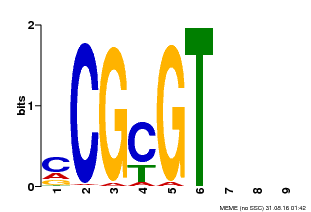
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011022796.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like isoform X1 | ||||
| Refseq | XP_011022797.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like isoform X1 | ||||
| Refseq | XP_011022798.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like isoform X1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | B9HFG2 | 0.0 | B9HFG2_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0007s05410.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8284 | 27 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



